<str<strong>on</strong>g>Proceedings</str<strong>on</strong>g> <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> <str<strong>on</strong>g>Third</str<strong>on</strong>g> <str<strong>on</strong>g>Internati<strong>on</strong>al</str<strong>on</strong>g> <str<strong>on</strong>g>C<strong>on</strong>ference</str<strong>on</strong>g> <strong>on</strong> <strong>Invasive</strong> SpartinaChapter 3: Ecosystem Effects <str<strong>on</strong>g>of</str<strong>on</strong>g> <strong>Invasive</strong> Spartinacorrecti<strong>on</strong> standards were IAEA S-1 (silver sulfide δ 34 S V-CDT= -0.3‰) and Iso-Analytical IA-R025 (barium sulfateδ 34 S V-CDT = +8.53‰). IA-R025 (barium sulfate δ 34 S V-CDT =+8.53‰) and whale baleen (δ 34 S V-CDT = +16.30‰) wereused as internal standards.The IsoSource isotopic mixing model analysisapplicati<strong>on</strong> (www.epa.gov/wed/pages/models.htm; Phillipsand Gregg 2003) was used to estimate <str<strong>on</strong>g>the</str<strong>on</strong>g> feasiblec<strong>on</strong>tributi<strong>on</strong>s <str<strong>on</strong>g>of</str<strong>on</strong>g> S. anglica and six o<str<strong>on</strong>g>the</str<strong>on</strong>g>r potential sources to<str<strong>on</strong>g>the</str<strong>on</strong>g> diets <str<strong>on</strong>g>of</str<strong>on</strong>g> Macoma, Mya, and Mytilus. IsoSource calculates<str<strong>on</strong>g>the</str<strong>on</strong>g> ranges <str<strong>on</strong>g>of</str<strong>on</strong>g> all possible source c<strong>on</strong>tributi<strong>on</strong>s when <str<strong>on</strong>g>the</str<strong>on</strong>g>number <str<strong>on</strong>g>of</str<strong>on</strong>g> sources placed in <str<strong>on</strong>g>the</str<strong>on</strong>g> mixing model exceeds n+1,where n is <str<strong>on</strong>g>the</str<strong>on</strong>g> number <str<strong>on</strong>g>of</str<strong>on</strong>g> isotopic tracers (e.g. δ 13 C or δ 34 S;Phillips and Gregg 2003). Therefore, IsoSource allowssource c<strong>on</strong>tributi<strong>on</strong>s to a mixture to be estimated regardless<str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> number <str<strong>on</strong>g>of</str<strong>on</strong>g> sources and isotopic tracers available. In<str<strong>on</strong>g>the</str<strong>on</strong>g>se circumstances unique soluti<strong>on</strong>s cannot be obtained dueto <str<strong>on</strong>g>the</str<strong>on</strong>g> use <str<strong>on</strong>g>of</str<strong>on</strong>g> large numbers <str<strong>on</strong>g>of</str<strong>on</strong>g> sources in <str<strong>on</strong>g>the</str<strong>on</strong>g> mixing model(Phillips and Gregg 2003). Since unique soluti<strong>on</strong>s cannot bedetermined, IsoSource output provides ranges <str<strong>on</strong>g>of</str<strong>on</strong>g> all possiblesource c<strong>on</strong>tributi<strong>on</strong>s to <str<strong>on</strong>g>the</str<strong>on</strong>g> mixture based <strong>on</strong> mass balancecalculati<strong>on</strong>s (Table 1; Phillips and Gregg 2003). The morenarrow <str<strong>on</strong>g>the</str<strong>on</strong>g> range <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g>se soluti<strong>on</strong>s (e.g 30%-50% vs. 10-70%), <str<strong>on</strong>g>the</str<strong>on</strong>g> more reliable <str<strong>on</strong>g>the</str<strong>on</strong>g> interpretati<strong>on</strong> <str<strong>on</strong>g>of</str<strong>on</strong>g> sourcec<strong>on</strong>tributi<strong>on</strong>s to <str<strong>on</strong>g>the</str<strong>on</strong>g> mixture can be. Thus, IsoSource is veryuseful for studies <str<strong>on</strong>g>of</str<strong>on</strong>g> trophic relati<strong>on</strong>ships where numerousfood sources could exist for a c<strong>on</strong>sumer. IsoSource has beenapplied to a variety <str<strong>on</strong>g>of</str<strong>on</strong>g> trophic studies inside and outside <str<strong>on</strong>g>of</str<strong>on</strong>g>estuaries (e.g. Felicetti et al. 2003; Melville and C<strong>on</strong>nolly2003; Ben-David et al. 2004; Newsome et al. 2004).The carb<strong>on</strong> isotope ratios <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> bivalves were adjustedby a 1‰ fracti<strong>on</strong>ati<strong>on</strong> factor based <strong>on</strong> Peters<strong>on</strong> and Fry(1987), Vander Zanden and Rasmussen (1999), andMcCutchan et al. (2003). Sulfur appears to have little to n<str<strong>on</strong>g>of</str<strong>on</strong>g>racti<strong>on</strong>ati<strong>on</strong> associated with its movement through trophiclevels. Due to <str<strong>on</strong>g>the</str<strong>on</strong>g> scarcity <str<strong>on</strong>g>of</str<strong>on</strong>g> sulfur fracti<strong>on</strong>ati<strong>on</strong> data(McCutchan et al. 2003), <str<strong>on</strong>g>the</str<strong>on</strong>g> typical assumpti<strong>on</strong> that <str<strong>on</strong>g>the</str<strong>on</strong>g>re isminimal fracti<strong>on</strong>ati<strong>on</strong> <str<strong>on</strong>g>of</str<strong>on</strong>g> sulfur was applied (Peters<strong>on</strong> andHowarth 1987). A biomass c<strong>on</strong>tributi<strong>on</strong> increment <str<strong>on</strong>g>of</str<strong>on</strong>g> 1%and a tolerance interval <str<strong>on</strong>g>of</str<strong>on</strong>g> 0.11‰ was used for all IsoSourceanalyses to insure that <str<strong>on</strong>g>the</str<strong>on</strong>g> full range <str<strong>on</strong>g>of</str<strong>on</strong>g> feasible soluti<strong>on</strong>swas generated (Table 1).RESULTSMacoma (n=12) clustered close to <str<strong>on</strong>g>the</str<strong>on</strong>g> outer edge <str<strong>on</strong>g>of</str<strong>on</strong>g>mixing polyg<strong>on</strong> in close proximity to living and deadSpartina tissue (Figure 1A). Only living tissue <str<strong>on</strong>g>of</str<strong>on</strong>g> Spartinawas c<strong>on</strong>sidered to be a comp<strong>on</strong>ent <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> diet <str<strong>on</strong>g>of</str<strong>on</strong>g> Macoma inall soluti<strong>on</strong>s <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> mixing model with c<strong>on</strong>tributi<strong>on</strong>s rangingfrom 37-60% (Table 1). Five <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> six productivity sourcessampled could be absent in <str<strong>on</strong>g>the</str<strong>on</strong>g> diet <str<strong>on</strong>g>of</str<strong>on</strong>g> Macoma (Table 1).Four Mya clustered near <str<strong>on</strong>g>the</str<strong>on</strong>g> center <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> mixingpolyg<strong>on</strong> while three o<str<strong>on</strong>g>the</str<strong>on</strong>g>r individuals clustered close toSpartina (Figure 1B). IsoSource indicated that Mya couldreceive 40-59% <str<strong>on</strong>g>of</str<strong>on</strong>g> its diet from living Spartina tissue andTable 1. IsoSource results. Ranges <str<strong>on</strong>g>of</str<strong>on</strong>g> feasible biomass source c<strong>on</strong>tributi<strong>on</strong>sfor <str<strong>on</strong>g>the</str<strong>on</strong>g> each bivalve summarized from 34,854 mass balanced mixingmodel soluti<strong>on</strong>s (Macoma balthica), 23,996 soluti<strong>on</strong>s (Mya arenaria), and56,083 soluti<strong>on</strong>s (Mytilus sp.). Each distributi<strong>on</strong> is defined by <str<strong>on</strong>g>the</str<strong>on</strong>g> minimumand maximum values as well as <str<strong>on</strong>g>the</str<strong>on</strong>g> 1 st and 99 th percentiles. The percentilesrepresent 98% <str<strong>on</strong>g>of</str<strong>on</strong>g> all <str<strong>on</strong>g>the</str<strong>on</strong>g> possible IsoSource soluti<strong>on</strong>s. The meanpercent <str<strong>on</strong>g>of</str<strong>on</strong>g> each source distributi<strong>on</strong> is cited to show central tendency <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g>distributi<strong>on</strong> <strong>on</strong>ly and should not be c<strong>on</strong>sidered a point estimate <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g>source c<strong>on</strong>tributi<strong>on</strong> to <str<strong>on</strong>g>the</str<strong>on</strong>g> diet <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> bivalve.Source C<strong>on</strong>sumer Biomass %Minimum 1% Mean 99% MaximumSpartina (living leaves) Macoma 37 41 52 59 60Mya 40 44 53 58 59Mytilus 19 24 36 43 44Spartina (standing dead) Macoma 0 0 15 38 46Mya 0 0 8 27 35Mytilus 0 0 11 36 46Distichlis spicata Macoma 0 0 10 26 31Mya 0 0 6 19 24Mytilus 0 0 8 25 32C 3 Vascular Plants Macoma 0 0 5 15 17Salicornia virginica Mya 3 5 14 23 26Grindelia integrifolia Mytilus 5 8 20 32 35Zostera marina Macoma 0 0 9 28 33Mya 0 0 10 32 40Mytilus 0 0 13 42 52Macroalgae Macoma 0 0 9 26 31Fucus spiralis Mya 0 0 9 30 38Mytilus 0 0 12 40 503-26% <str<strong>on</strong>g>of</str<strong>on</strong>g> its diet from C 3 vegetati<strong>on</strong>. All o<str<strong>on</strong>g>the</str<strong>on</strong>g>r sources hadwide ranges <str<strong>on</strong>g>of</str<strong>on</strong>g> potential producer c<strong>on</strong>tributi<strong>on</strong>s that includedzero as likely dietary c<strong>on</strong>tributi<strong>on</strong>s (Table 1).Living Spartina biomass and C 3 vegetati<strong>on</strong> were <str<strong>on</strong>g>the</str<strong>on</strong>g><strong>on</strong>ly dietary sources that IsoSource included within all <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g>feasible soluti<strong>on</strong>s for Mytilus. Living Spartina couldc<strong>on</strong>tribute 19-44% to <str<strong>on</strong>g>the</str<strong>on</strong>g> diet <str<strong>on</strong>g>of</str<strong>on</strong>g> Mytilus, whereas C 3vegetati<strong>on</strong> may c<strong>on</strong>tribute 5-35% <str<strong>on</strong>g>of</str<strong>on</strong>g> its diet. All o<str<strong>on</strong>g>the</str<strong>on</strong>g>rpotential sources included 0% as <str<strong>on</strong>g>the</str<strong>on</strong>g> most frequent feasiblec<strong>on</strong>tributi<strong>on</strong> (Table 1). Six <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> seven individuals <str<strong>on</strong>g>of</str<strong>on</strong>g>Mytilus clustered in <str<strong>on</strong>g>the</str<strong>on</strong>g> center <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> mixing polyg<strong>on</strong> while<strong>on</strong>e individual was more depleted in its δ 34 S signature(Fig. 1C).DISCUSSIONOur mixing models present evidence that Spartinaanglica is c<strong>on</strong>sumed by bivalves. The use <str<strong>on</strong>g>of</str<strong>on</strong>g> Spartina byMacoma is particularly c<strong>on</strong>vincing based <strong>on</strong> <str<strong>on</strong>g>the</str<strong>on</strong>g> c<strong>on</strong>strainedmixing model soluti<strong>on</strong> and <str<strong>on</strong>g>the</str<strong>on</strong>g> proximity <str<strong>on</strong>g>of</str<strong>on</strong>g> individuals to<str<strong>on</strong>g>the</str<strong>on</strong>g> isotopic signatures <str<strong>on</strong>g>of</str<strong>on</strong>g> living and dead Spartina (Fig. 1A).The mixing model for Mya is also c<strong>on</strong>vincing, although <str<strong>on</strong>g>the</str<strong>on</strong>g>δ 13 C values tend to be more depleted than those <str<strong>on</strong>g>of</str<strong>on</strong>g> Macomapotentially indicating smaller Spartina c<strong>on</strong>tributi<strong>on</strong>s to itsdiet compared to Macoma. With <str<strong>on</strong>g>the</str<strong>on</strong>g> excepti<strong>on</strong> <str<strong>on</strong>g>of</str<strong>on</strong>g> <strong>on</strong>eoutlying individual, Mytilus had <str<strong>on</strong>g>the</str<strong>on</strong>g> most c<strong>on</strong>sistent isotopicsignatures for δ 13 C and δ 34 S, and <str<strong>on</strong>g>the</str<strong>on</strong>g> lowest ranges <str<strong>on</strong>g>of</str<strong>on</strong>g>potential Spartina c<strong>on</strong>tributi<strong>on</strong>s. As a suspensi<strong>on</strong> feedingmussel, Mytilus should be drawing its food from <str<strong>on</strong>g>the</str<strong>on</strong>g>- 155 -
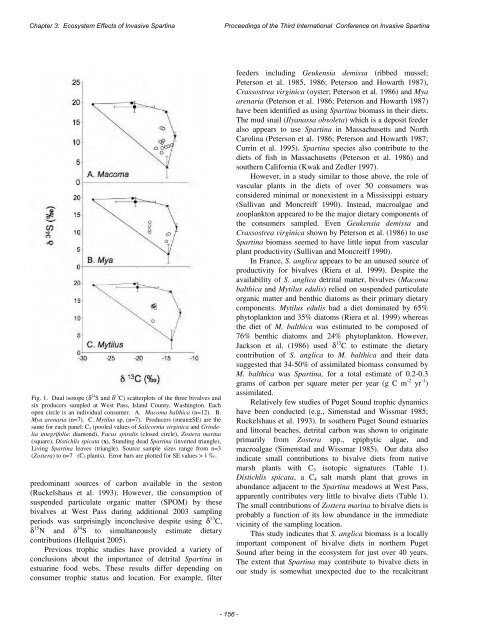
Chapter 3: Ecosystem Effects <str<strong>on</strong>g>of</str<strong>on</strong>g> <strong>Invasive</strong> Spartina<str<strong>on</strong>g>Proceedings</str<strong>on</strong>g> <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> <str<strong>on</strong>g>Third</str<strong>on</strong>g> <str<strong>on</strong>g>Internati<strong>on</strong>al</str<strong>on</strong>g> <str<strong>on</strong>g>C<strong>on</strong>ference</str<strong>on</strong>g> <strong>on</strong> <strong>Invasive</strong> SpartinaFig. 1. Dual isotope (δ 34 Sand δ 13 C) scatterplots <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> three bivalves andsix producers sampled at West Pass, Island County, Washingt<strong>on</strong>. Eachopen circle is an individual c<strong>on</strong>sumer. A. Macoma balthica (n=12). B.Mya arenaria (n=7). C. Mytilus sp. (n=7). Producers (mean±SE) are <str<strong>on</strong>g>the</str<strong>on</strong>g>same for each panel: C 3 (pooled values <str<strong>on</strong>g>of</str<strong>on</strong>g> Salicornia virginica and Grindeliaintegrifolia: diam<strong>on</strong>d), Fucus spiralis (closed circle), Zostera marina(square), Distichlis spicata (x), Standing dead Spartina (inverted triangle),Living Spartina leaves (triangle). Source sample sizes range from n=3(Zostera) to n=7 (C 3 plants). Error bars are plotted for SE values > 1 ‰.predominant sources <str<strong>on</strong>g>of</str<strong>on</strong>g> carb<strong>on</strong> available in <str<strong>on</strong>g>the</str<strong>on</strong>g> sest<strong>on</strong>(Ruckelshaus et al. 1993). However, <str<strong>on</strong>g>the</str<strong>on</strong>g> c<strong>on</strong>sumpti<strong>on</strong> <str<strong>on</strong>g>of</str<strong>on</strong>g>suspended particulate organic matter (SPOM) by <str<strong>on</strong>g>the</str<strong>on</strong>g>sebivalves at West Pass during additi<strong>on</strong>al 2003 samplingperiods was surprisingly inc<strong>on</strong>clusive despite using δ 13 C,δ 15 N and δ 34 S to simultaneously estimate dietaryc<strong>on</strong>tributi<strong>on</strong>s (Hellquist 2005).Previous trophic studies have provided a variety <str<strong>on</strong>g>of</str<strong>on</strong>g>c<strong>on</strong>clusi<strong>on</strong>s about <str<strong>on</strong>g>the</str<strong>on</strong>g> importance <str<strong>on</strong>g>of</str<strong>on</strong>g> detrital Spartina inestuarine food webs. These results differ depending <strong>on</strong>c<strong>on</strong>sumer trophic status and locati<strong>on</strong>. For example, filterfeeders including Geukensia demissa (ribbed mussel;Peters<strong>on</strong> et al. 1985, 1986; Peters<strong>on</strong> and Howarth 1987),Crassostrea virginica (oyster; Peters<strong>on</strong> et al. 1986) and Myaarenaria (Peters<strong>on</strong> et al. 1986; Peters<strong>on</strong> and Howarth 1987)have been identified as using Spartina biomass in <str<strong>on</strong>g>the</str<strong>on</strong>g>ir diets.The mud snail (Ilyanassa obsoleta) which is a deposit feederalso appears to use Spartina in Massachusetts and NorthCarolina (Peters<strong>on</strong> et al. 1986; Peters<strong>on</strong> and Howarth 1987;Currin et al. 1995). Spartina species also c<strong>on</strong>tribute to <str<strong>on</strong>g>the</str<strong>on</strong>g>diets <str<strong>on</strong>g>of</str<strong>on</strong>g> fish in Massachusetts (Peters<strong>on</strong> et al. 1986) andsou<str<strong>on</strong>g>the</str<strong>on</strong>g>rn California (Kwak and Zedler 1997).However, in a study similar to those above, <str<strong>on</strong>g>the</str<strong>on</strong>g> role <str<strong>on</strong>g>of</str<strong>on</strong>g>vascular plants in <str<strong>on</strong>g>the</str<strong>on</strong>g> diets <str<strong>on</strong>g>of</str<strong>on</strong>g> over 50 c<strong>on</strong>sumers wasc<strong>on</strong>sidered minimal or n<strong>on</strong>existent in a Mississippi estuary(Sullivan and M<strong>on</strong>creiff 1990). Instead, macroalgae andzooplankt<strong>on</strong> appeared to be <str<strong>on</strong>g>the</str<strong>on</strong>g> major dietary comp<strong>on</strong>ents <str<strong>on</strong>g>of</str<strong>on</strong>g><str<strong>on</strong>g>the</str<strong>on</strong>g> c<strong>on</strong>sumers sampled. Even Geukensia demissa andCrassostrea virginica shown by Peters<strong>on</strong> et al. (1986) to useSpartina biomass seemed to have little input from vascularplant productivity (Sullivan and M<strong>on</strong>creiff 1990).In France, S. anglica appears to be an unused source <str<strong>on</strong>g>of</str<strong>on</strong>g>productivity for bivalves (Riera et al. 1999). Despite <str<strong>on</strong>g>the</str<strong>on</strong>g>availability <str<strong>on</strong>g>of</str<strong>on</strong>g> S. anglica detrital matter, bivalves (Macomabalthica and Mytilus edulis) relied <strong>on</strong> suspended particulateorganic matter and benthic diatoms as <str<strong>on</strong>g>the</str<strong>on</strong>g>ir primary dietarycomp<strong>on</strong>ents. Mytilus edulis had a diet dominated by 65%phytoplankt<strong>on</strong> and 35% diatoms (Riera et al. 1999) whereas<str<strong>on</strong>g>the</str<strong>on</strong>g> diet <str<strong>on</strong>g>of</str<strong>on</strong>g> M. balthica was estimated to be composed <str<strong>on</strong>g>of</str<strong>on</strong>g>76% benthic diatoms and 24% phytoplankt<strong>on</strong>. However,Jacks<strong>on</strong> et al. (1986) used δ 13 C to estimate <str<strong>on</strong>g>the</str<strong>on</strong>g> dietaryc<strong>on</strong>tributi<strong>on</strong> <str<strong>on</strong>g>of</str<strong>on</strong>g> S. anglica to M. balthica and <str<strong>on</strong>g>the</str<strong>on</strong>g>ir datasuggested that 34-50% <str<strong>on</strong>g>of</str<strong>on</strong>g> assimilated biomass c<strong>on</strong>sumed byM. balthica was Spartina, for a total estimate <str<strong>on</strong>g>of</str<strong>on</strong>g> 0.2-0.3grams <str<strong>on</strong>g>of</str<strong>on</strong>g> carb<strong>on</strong> per square meter per year (g C m -2 yr -1 )assimilated.Relatively few studies <str<strong>on</strong>g>of</str<strong>on</strong>g> Puget Sound trophic dynamicshave been c<strong>on</strong>ducted (e.g., Simenstad and Wissmar 1985;Ruckelshaus et al. 1993). In sou<str<strong>on</strong>g>the</str<strong>on</strong>g>rn Puget Sound estuariesand littoral beaches, detrital carb<strong>on</strong> was shown to originateprimarily from Zostera spp., epiphytic algae, andmacroalgae (Simenstad and Wissmar 1985). Our data alsoindicate small c<strong>on</strong>tributi<strong>on</strong>s to bivalve diets from nativemarsh plants with C 3 isotopic signatures (Table 1).Distichlis spicata, a C 4 salt marsh plant that grows inabundance adjacent to <str<strong>on</strong>g>the</str<strong>on</strong>g> Spartina meadows at West Pass,apparently c<strong>on</strong>tributes very little to bivalve diets (Table 1).The small c<strong>on</strong>tributi<strong>on</strong>s <str<strong>on</strong>g>of</str<strong>on</strong>g> Zostera marina to bivalve diets isprobably a functi<strong>on</strong> <str<strong>on</strong>g>of</str<strong>on</strong>g> its low abundance in <str<strong>on</strong>g>the</str<strong>on</strong>g> immediatevicinity <str<strong>on</strong>g>of</str<strong>on</strong>g> <str<strong>on</strong>g>the</str<strong>on</strong>g> sampling locati<strong>on</strong>.This study indicates that S. anglica biomass is a locallyimportant comp<strong>on</strong>ent <str<strong>on</strong>g>of</str<strong>on</strong>g> bivalve diets in nor<str<strong>on</strong>g>the</str<strong>on</strong>g>rn PugetSound after being in <str<strong>on</strong>g>the</str<strong>on</strong>g> ecosystem for just over 40 years.The extent that Spartina may c<strong>on</strong>tribute to bivalve diets inour study is somewhat unexpected due to <str<strong>on</strong>g>the</str<strong>on</strong>g> recalcitrant- 156 -
- Page 2 and 3:
Proceedings <stron
- Page 4 and 5:
FORWARD & ACKNOWLEDGEMENTSThe <stro
- Page 6 and 7:
TABLE OF CONTENTSForward & Acknowle
- Page 9 and 10:
Community Spartina Education and St
- Page 11 and 12:
included the docum
- Page 14:
CHAPTER ONESpartina Biology
- Page 17 and 18:
Chapter 1: Spartina Biology
- Page 19 and 20:
Chapter 1: Spartina Biology
- Page 21 and 22:
Chapter 1: Spartina Biology
- Page 23 and 24:
Chapter 1: Spartina Biology
- Page 25 and 26:
Chapter 1: Spartina Biology
- Page 28 and 29:
Proceedings <stron
- Page 30 and 31:
Proceedings <stron
- Page 32 and 33:
Proceedings <stron
- Page 34:
Proceedings <stron
- Page 37 and 38:
Chapter 1: Spartina Biology
- Page 39 and 40:
Chapter 1: Spartina Biology
- Page 42 and 43:
Proceedings <stron
- Page 44:
Proceedings <stron
- Page 47 and 48:
Chapter 1: Spartina Biology
- Page 49 and 50:
Chapter 1: Spartina Biology
- Page 51 and 52:
Chapter 1: Spartina Biology
- Page 53 and 54:
Chapter 1: Spartina Biology
- Page 55 and 56:
Chapter 1: Spartina Biology
- Page 57 and 58:
Chapter 1: Spartina Biology
- Page 60 and 61:
Proceedings <stron
- Page 62 and 63:
Proceedings <stron
- Page 64 and 65:
Proceedings <stron
- Page 66:
Proceedings <stron
- Page 69 and 70:
Chapter 1: Spartina Biology
- Page 71 and 72:
Chapter 1: Spartina Biology
- Page 74 and 75:
Proceedings <stron
- Page 76:
Proceedings <stron
- Page 79 and 80:
Chapter 2: Spartina Distribution an
- Page 81 and 82:
Chapter 2: Spartina Distribution an
- Page 83 and 84:
Chapter 2: Spartina Distribution an
- Page 86 and 87:
Proceedings <stron
- Page 88 and 89:
Proceedings <stron
- Page 90 and 91:
Proceedings <stron
- Page 92 and 93:
Proceedings <stron
- Page 94 and 95:
Proceedings <stron
- Page 96 and 97:
Proceedings <stron
- Page 98:
Proceedings <stron
- Page 101 and 102:
Chapter 2: Spartina Distribution an
- Page 103 and 104:
Chapter 2: Spartina Distribution an
- Page 105 and 106:
Chapter 2: Spartina Distribution an
- Page 108 and 109:
Proceedings <stron
- Page 110:
Proceedings <stron
- Page 113 and 114:
Chapter 2: Spartina Distribution an
- Page 115 and 116:
Chapter 2: Spartina Distribution an
- Page 117 and 118: Chapter 2: Spartina Distribution an
- Page 119 and 120: Chapter 2: Spartina Distribution an
- Page 122 and 123: Proceedings <stron
- Page 124 and 125: Proceedings <stron
- Page 126 and 127: Proceedings <stron
- Page 128: Proceedings <stron
- Page 131 and 132: Chapter 2: Spartina Distribution an
- Page 134 and 135: Proceedings <stron
- Page 136 and 137: Proceedings <stron
- Page 138 and 139: Proceedings <stron
- Page 140: CHAPTER THREEEcosystem Effects <str
- Page 143 and 144: Chapter 3: Ecosystem Effects <stron
- Page 145 and 146: Chapter 3: Ecosystem Effects <stron
- Page 148 and 149: Proceedings <stron
- Page 150 and 151: Proceedings <stron
- Page 152: Proceedings <stron
- Page 155 and 156: Chapter 3: Ecosystem Effects <stron
- Page 157 and 158: Chapter 3: Ecosystem Effects <stron
- Page 160 and 161: Proceedings <stron
- Page 162 and 163: Proceedings <stron
- Page 164: Proceedings <stron
- Page 167: Chapter 3: Ecosystem Effects <stron
- Page 171 and 172: Chapter 3: Ecosystem Effects <stron
- Page 174 and 175: Proceedings <stron
- Page 176: Proceedings <stron
- Page 179 and 180: Chapter 3: Ecosystem Effects <stron
- Page 181 and 182: Chapter 3: Ecosystem Effects <stron
- Page 184 and 185: Proceedings <stron
- Page 186 and 187: Proceedings <stron
- Page 188 and 189: Proceedings <stron
- Page 190 and 191: Proceedings <stron
- Page 192 and 193: Proceedings <stron
- Page 194 and 195: Proceedings <stron
- Page 196: Proceedings <stron
- Page 199 and 200: Chapter 3: Ecosystem Effects <stron
- Page 201 and 202: Chapter 3: Ecosystem Effects <stron
- Page 204 and 205: Proceedings <stron
- Page 206 and 207: Proceedings <stron
- Page 208 and 209: Proceedings <stron
- Page 210 and 211: Proceedings <stron
- Page 212: Proceedings <stron
- Page 216 and 217: Proceedings <stron
- Page 218 and 219:
Proceedings <stron
- Page 220 and 221:
Proceedings <stron
- Page 222 and 223:
Proceedings <stron
- Page 224 and 225:
Proceedings <stron
- Page 226 and 227:
Proceedings <stron
- Page 228 and 229:
Proceedings <stron
- Page 230 and 231:
Proceedings <stron
- Page 232 and 233:
Proceedings <stron
- Page 234 and 235:
Proceedings <stron
- Page 236 and 237:
Proceedings <stron
- Page 238 and 239:
Proceedings <stron
- Page 240 and 241:
Proceedings <stron
- Page 242 and 243:
Proceedings <stron
- Page 244 and 245:
Proceedings <stron
- Page 246:
Proceedings <stron
- Page 249 and 250:
Chapter 4: Spartina Control and Man
- Page 251 and 252:
Chapter 4: Spartina Control and Man
- Page 253 and 254:
Chapter 4: Spartina Control and Man
- Page 255 and 256:
Chapter 4: Spartina Control and Man
- Page 257 and 258:
Chapter 4: Spartina Control and Man
- Page 259 and 260:
Chapter 4: Spartina Control and Man
- Page 261 and 262:
Chapter 4: Spartina Control and Man
- Page 263 and 264:
Chapter 4: Spartina Control and Man
- Page 265 and 266:
Chapter 4: Spartina Control and Man
- Page 267 and 268:
Chapter 4: Spartina Control and Man
- Page 269 and 270:
Chapter 4: Spartina Control and Man
- Page 271 and 272:
Chapter 4: Spartina Control and Man
- Page 273 and 274:
Chapter 4: Spartina Control and Man
- Page 276 and 277:
Proceedings <stron
- Page 278 and 279:
Proceedings <stron
- Page 280 and 281:
Proceedings <stron
- Page 282 and 283:
Proceedings <stron
- Page 284 and 285:
Proceedings <stron
- Page 286 and 287:
Proceedings <stron
- Page 288 and 289:
Proceedings <stron
- Page 290:
Proceedings <stron