Segmentation of Stochastic Images using ... - Jacobs University
Segmentation of Stochastic Images using ... - Jacobs University
Segmentation of Stochastic Images using ... - Jacobs University
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
Chapter 7 <strong>Stochastic</strong> Level Sets<br />
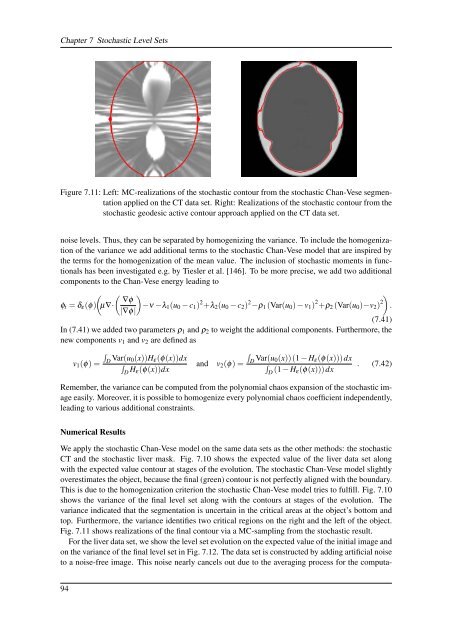
Figure 7.11: Left: MC-realizations <strong>of</strong> the stochastic contour from the stochastic Chan-Vese segmentation<br />
applied on the CT data set. Right: Realizations <strong>of</strong> the stochastic contour from the<br />
stochastic geodesic active contour approach applied on the CT data set.<br />
noise levels. Thus, they can be separated by homogenizing the variance. To include the homogenization<br />
<strong>of</strong> the variance we add additional terms to the stochastic Chan-Vese model that are inspired by<br />
the terms for the homogenization <strong>of</strong> the mean value. The inclusion <strong>of</strong> stochastic moments in functionals<br />
has been investigated e.g. by Tiesler et al. [146]. To be more precise, we add two additional<br />
components to the Chan-Vese energy leading to<br />
( ( )<br />
∇φ<br />
φ t = δ ε (φ) µ∇·<br />
)−ν −λ 1 (u 0 − c 1 ) 2 +λ 2 (u 0 − c 2 ) 2 −ρ 1 (Var(u 0 ) − v 1 ) 2 +ρ 2 (Var(u 0 )−v 2 ) 2 .<br />
|∇φ|<br />
(7.41)<br />
In (7.41) we added two parameters ρ 1 and ρ 2 to weight the additional components. Furthermore, the<br />
new components v 1 and v 2 are defined as<br />
∫<br />
D<br />
v 1 (φ) =<br />
Var(u 0(x))H ε (φ(x))dx<br />
∫<br />
D H ε(φ(x))dx<br />
∫<br />
and v 2 (φ) =<br />
D Var(u 0(x))(1 − H ε (φ(x)))dx<br />
∫<br />
D (1 − H ε(φ(x)))dx<br />
. (7.42)<br />
Remember, the variance can be computed from the polynomial chaos expansion <strong>of</strong> the stochastic image<br />
easily. Moreover, it is possible to homogenize every polynomial chaos coefficient independently,<br />
leading to various additional constraints.<br />
Numerical Results<br />
We apply the stochastic Chan-Vese model on the same data sets as the other methods: the stochastic<br />
CT and the stochastic liver mask. Fig. 7.10 shows the expected value <strong>of</strong> the liver data set along<br />
with the expected value contour at stages <strong>of</strong> the evolution. The stochastic Chan-Vese model slightly<br />
overestimates the object, because the final (green) contour is not perfectly aligned with the boundary.<br />
This is due to the homogenization criterion the stochastic Chan-Vese model tries to fulfill. Fig. 7.10<br />
shows the variance <strong>of</strong> the final level set along with the contours at stages <strong>of</strong> the evolution. The<br />
variance indicated that the segmentation is uncertain in the critical areas at the object’s bottom and<br />
top. Furthermore, the variance identifies two critical regions on the right and the left <strong>of</strong> the object.<br />
Fig. 7.11 shows realizations <strong>of</strong> the final contour via a MC-sampling from the stochastic result.<br />
For the liver data set, we show the level set evolution on the expected value <strong>of</strong> the initial image and<br />
on the variance <strong>of</strong> the final level set in Fig. 7.12. The data set is constructed by adding artificial noise<br />
to a noise-free image. This noise nearly cancels out due to the averaging process for the computa-<br />
94