Segmentation of Stochastic Images using ... - Jacobs University
Segmentation of Stochastic Images using ... - Jacobs University
Segmentation of Stochastic Images using ... - Jacobs University
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
8.1 Random Walker <strong>Segmentation</strong> with <strong>Stochastic</strong> Parameter<br />
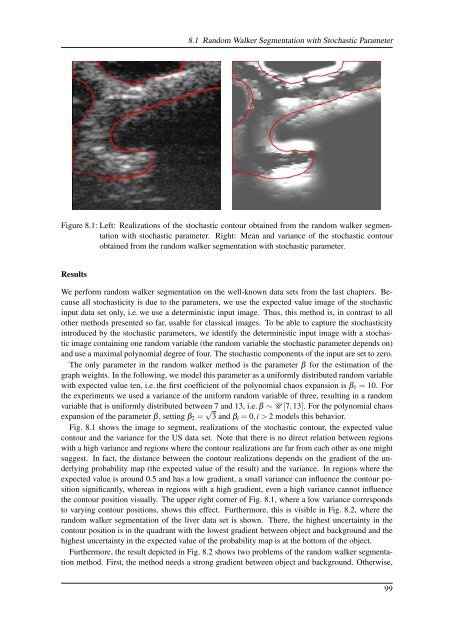
Figure 8.1: Left: Realizations <strong>of</strong> the stochastic contour obtained from the random walker segmentation<br />
with stochastic parameter. Right: Mean and variance <strong>of</strong> the stochastic contour<br />
obtained from the random walker segmentation with stochastic parameter.<br />
Results<br />
We perform random walker segmentation on the well-known data sets from the last chapters. Because<br />
all stochasticity is due to the parameters, we use the expected value image <strong>of</strong> the stochastic<br />
input data set only, i.e. we use a deterministic input image. Thus, this method is, in contrast to all<br />
other methods presented so far, usable for classical images. To be able to capture the stochasticity<br />
introduced by the stochastic parameters, we identify the deterministic input image with a stochastic<br />
image containing one random variable (the random variable the stochastic parameter depends on)<br />
and use a maximal polynomial degree <strong>of</strong> four. The stochastic components <strong>of</strong> the input are set to zero.<br />
The only parameter in the random walker method is the parameter β for the estimation <strong>of</strong> the<br />
graph weights. In the following, we model this parameter as a uniformly distributed random variable<br />
with expected value ten, i.e. the first coefficient <strong>of</strong> the polynomial chaos expansion is β 1 = 10. For<br />
the experiments we used a variance <strong>of</strong> the uniform random variable <strong>of</strong> three, resulting in a random<br />
variable that is uniformly distributed between 7 and 13, i.e. β ∼ U [7,13]. For the polynomial chaos<br />
expansion <strong>of</strong> the parameter β, setting β 2 = √ 3 and β i = 0,i > 2 models this behavior.<br />
Fig. 8.1 shows the image to segment, realizations <strong>of</strong> the stochastic contour, the expected value<br />
contour and the variance for the US data set. Note that there is no direct relation between regions<br />
with a high variance and regions where the contour realizations are far from each other as one might<br />
suggest. In fact, the distance between the contour realizations depends on the gradient <strong>of</strong> the underlying<br />
probability map (the expected value <strong>of</strong> the result) and the variance. In regions where the<br />
expected value is around 0.5 and has a low gradient, a small variance can influence the contour position<br />
significantly, whereas in regions with a high gradient, even a high variance cannot influence<br />
the contour position visually. The upper right corner <strong>of</strong> Fig. 8.1, where a low variance corresponds<br />
to varying contour positions, shows this effect. Furthermore, this is visible in Fig. 8.2, where the<br />
random walker segmentation <strong>of</strong> the liver data set is shown. There, the highest uncertainty in the<br />
contour position is in the quadrant with the lowest gradient between object and background and the<br />
highest uncertainty in the expected value <strong>of</strong> the probability map is at the bottom <strong>of</strong> the object.<br />
Furthermore, the result depicted in Fig. 8.2 shows two problems <strong>of</strong> the random walker segmentation<br />
method. First, the method needs a strong gradient between object and background. Otherwise,<br />
99