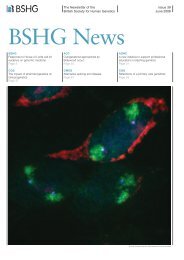
spliceosome. They have now focused on generating phosphor-specific antibodies in order to visualize catalytically activespliceosomes in the cell nucleus. For example, SF3b155 is a U2 snRNP specific protein that is specifically phosphorylatedonly upon spliceosome activation and its dephosphorylation is required for the second step of splicing. PhosphorylatedSF3b155 is thus specific for active spliceosomes. To visualize these by immunofluorescence (IF) in HeLa cells, theygenerated phospho-peptide specific antibodies (R79) specifically recognizing SF3b155 phosphorylation sites, which werefirst determined by mass spectrometry (collaboration with H. Urlaub). The antibodies were highly specific forphosphorylated SF3b155 on Western blots. IF studies with HeLa cells revealed a punctated staining pattern exclusively inthe nucleoplasm. Extensive further double IF studies revealed that the R79 signal i) was strictly dependent on ongoingtranscription, ii) did not overlap with nuclear speckles, and thus iii) constituted only a subset of all U2 snRNP containingregions within the nucleoplasm. Interestingly, the R79 signal co-localized with sites of newly synthesized RNA, consistentwith the idea of co-transcriptional splicing, but also at non-overlapping sites. Consistent with this observation, the R79 signalpartially overlapped with actively transcribing RNA polymerase II. Taken together, the data suggest a unique localization ofcatalytically active spliceosomes in the cell nucleus.200 PP1 regulation of the interaction of RNA with the splicing factos SF2/ASF, tra2-beta1 and SRp30c.Group 3 (Stamm) has previously shown that the phosphatase PP1 binds to the RRM of several splicing factors and regulatesalternative pre-mRNA processing. In the period of this report, this group showed that PP1 has two functions when binding tothe RRM of splicing factors. First, it dephosphorylates splicing factors as previously reported. The second function that isnow under investigation is that PP1 causes the release of RNA from the RRM.201 Relationship between B52 and the other candidate genes.A genetic screen performed by Group 12a (Tazi) resulted in the identification of DNA topoisomerase I (Topo I) as asuppressor of the developmental defects caused by overexpression of the Drosophila SR protein B52 in vivo. Thus, thisestablished an unsuspected tight coupling between the splicing factor B52 and Topo I during Drosophila development. Thisgroup now found that the localization of Topo I perfectly matches the localization of B52 on polytene chromosome at allloci. In transgenic flies, expression of a high affinity binding site for B52 restricted localization of both B52 and Topo I tothis single transcription site, whereas B52 RNAi knockdown induced mis-localization of Topo I in the nucleolus (Juge et al.,submitted). Unexpectedly, specific overexpression of B52 in salivary glands allowed massive recruitment of both B52 andTopo I, but not RNA Pol II, on nuclear RNA molecules, demonstrating for the first time that Topo I may assemble on RNPsto coordinate the action of SR proteins during transcription. This coordination likely involves an intrinsic kinase activity ofTopo I, which was shown to be required to specifically phosphorylate B52 in vivo. Thus, the partnership between Topo I andB52 for the specific phosphorylation of serine residues located within the RS domain and the control of DNA supercoïlingcould be key determinants for functional integration of transcription and RNA processing machineries in a mutuallybeneficial manner for efficient and regulated gene expression.202 Role of Akt as a novel SR protein kinase.Finally, Group 20b (Srebrow), a member YIP program, has continued her study on the regulation of Rac1 pre-mRNAalternative splicing in a model of epithelial-mesenchymal transition induced by matrix metalloproteases, in particular MMP-3 (stromelysin-1). They had already identified hnRNPA1 and hnRNP A2 as two regulatory proteins that were able to bindRac1 alternative exon 3b. They constructed an exon 3b splicing reporter minigene and used a set of biochemical assays suchas RNA-affinity purification, UV-cross linking followed by immunoprecipitation, and RNA gel shift, to start deciphering themechanism by which MMP-3 causes an increase in exon 3b inclusion and to further characterize the involvement of hnRNPA1. They defined a sequence element within exon 3b that resembles the hnRNP A1 consensus binding site. Mutation of thiselement results in a decreased binding of hnRNP A1 to this exon compared to the wild-type one. Furthermore, overexpressionof hnRNPA1 inhibits 3b exon inclusion in transcripts derived from the Rac1 3b minigene. Interestingly, theyfound that there is less hnRNP A1-exon 3b complex formation with nuclear extracts from MMP-3 treated cells than withnuclear extracts from untreated ones. These and other results from their lab suggest that other regulatory factors may becooperating with hnRNP A1 to repress inclusion of 3b exon under basal conditions and to induce its inclusion upontreatment with MMP-3. While exploring post-translational modifications of hnRNP A1 that could be involved in regulatingits activity upon MMP-3 treatment, they started working in the SUMOylation pathway and obtained unexpected resultssuggesting that certain SR proteins may influence this process.Problems and explanations for delays, postponements etc.202 Srebrow has encountered technical difficulties related to the growth and responsiveness of the cell culture system use tostudy Rac1 splicing regulation and they have spent considerable time and effort to overcome these problems. This hasresulted in lack of progress with respect to deliverable 202, the role of Akt as an SR protein kinase. This is now presented asa deliverable for the next period.PUBLICATIONSAkusjarvi,G. (<strong>2008</strong>) Temporal regulation of adenovirus major late alternative RNA splicing. Front. Biosci., 13, 5006-5015.Bessonov,S., Anokhina,M., Will,C.L., Urlaub,H. and Lührmann,R. (<strong>2008</strong>)49
Isolation of an active step I spliceosome and composition of its RNP core. Nature, 452, 846-850.Biamonti,G. and Cáceres,J.F. (2009) Cellular stress and RNA splicing. Trends Biochem..Sci., in press.Ellis,J.D., Lleres,D., Denegri,M., Lamond,A.I. and Cáceres,J.F. (<strong>2008</strong>) Spatial mapping of splicing factor complexesinvolved in exon and intron definition. J.Cell. Biol., 181, 921-934Comment/ Reviews on this paperRobinson,R. (<strong>2008</strong>) A better way to see splicing partners. J. Cell Biol., 181, 875.Grosso,A.R., Martins,S. and Carmo-Fonseca,M. (<strong>2008</strong>) The emerging role of splicing factors in cancer. EMBO Rep., 9,1087-1093.Herold,N., Will,C.L., Wolf,E., Kastner,B., Urlaub,H. and Lührmann,R. (2009) Conservation of the protein compositionand electron microscopy structure of Drosophila melanogaster and human spliceosomal complexes. Mol Cell Biol., 29,281-301.Kuhn,A.N., van Santen,M.A., Schwienhorst,A., Urlaub,H. and Lührmann,R. (2009) Stalling of spliceosome assembly atdistinct stages by small-molecule inhibitors of protein acetylation and deacetylation. RNA, 15, 153-175.Mathew,R., Hartmuth,K., Mohlmann,S., Urlaub,H., Ficner,R. and Lührmann,R. (<strong>2008</strong>) Phosphorylation of human PRP28by SRPK2 is required for integration of the U4/U6-U5 tri-snRNP into the spliceosome. Nat. Struct. Mol. Biol., 15, 435-43.Comment/ Reviews on this paperMaeder,C. and Guthrie,C. (<strong>2008</strong>) Modifications target spliceosome dynamics. Nat. Struct. Mol. Biol., 15, 426-428.Michlewski,G., Sanford,J.R. and Cáceres,J.F. (<strong>2008</strong>) The splicing factor SF2/ASF regulates translation initiation byenhancing phosphorylation of 4E-BP1. Mol Cell., 30, 179-189.Comment/ Reviews on this paperBushell,M., Stoneley,M., Spriggs,K.A. and Willis,A.E. (<strong>2008</strong>) SF2/ASF TORCs Up Translation. Mol Cell, 30, 262-263Rino,J., Desterro, J.M., Pacheco,T.R., Gadella,T.W. and Carmo-Fonseca,M.A. (<strong>2008</strong>) Splicing factors SF1 and U2AFassociate in extra-spliceosomal complexes. Mol.Cell.Biol., 28, 3045-57.Sanford,J.R., Coutinho,P., Hacket, J.A., Wang,X., Ranahan,W. and Cáceres,J.F. (<strong>2008</strong>) Identification of nuclear andcytoplasmic mRNA targets for the shuttling protein SF2/ASF. PLoS ONE, 3(10): e3369.Stamm,S. (<strong>2008</strong>) Regulation of alternative splicing by reversible phosphorylation. J.Biol. Chem., 283, 1223-1227.Sumanasekera, C., Watt,D.S. and Stamm,S. (<strong>2008</strong>) Substances that can changealternative splice-site selection. Biochem. Soc. Trans., 36, 483-490.Tazi,J., Bakkour,N. and Stamm,S. (2009) Alternative splicing and disease. Biochim Biophys Acta. 1792, 14-26.Valgardsdottir,R., Chiodi,I., Giordano,M., Rossi,A., Bazzini,S., Ghigna,C., Riva,S. and Biamonti,G. (<strong>2008</strong>) Transcription ofSatellite III non-coding RNAs is a general stress response in human cells. Nucleic Acids Res., 36, 423-434.Yu,Y., Maroney,P.A., Denker,J.A., Zhang,X.H., Dybkov,O., Lührmann,R., Jankowsky,E., Chasin,L.A. and Nilsen,T.W.(<strong>2008</strong>) Dynamic regulation of alternative splicing by silencers that modulate 5' splice site competition. Cell, 135, 1224-1236.MANUSCRIPTSBackstrom,E., Kaufmann,K. and Akusjarvi,G. The adenovirus L4-22K protein is a keyviral transcriptional regulator. Manuscript in preparation.Juge,F., Fic,W, Fernando,C. and Tazi,J. Coordinated action of the splicing factor B52 and DNA topoisomerase I inDrosophila development. submitted for publication.Tuduri.S, Crabbe,L., Tourriere,H., Conti,C., Jauch,A., Pantesco,V., de Vos,J., Theillet,C., Pommier,Y., Tazi,J., Coquelle,A.and Pasero,P. Topoisomerase I suppresses genomic instability by preventing interference between transcription and DNAreplication. Nat. Cell Biol., under revision.50
- Page 3 and 4: TABLE OF CONTENTSA. PERIODIC ACTIVI
- Page 5 and 6: 1 PUBLISHABLE EXECUTIVE SUMMARY EUR
- Page 7 and 8: Dr. Davide Gabellini(28) Swiss Inst
- Page 9 and 10: WP7 Molecular characterization of s
- Page 11 and 12: To identify protein-protein interac
- Page 13 and 14: expressed in a particular tissue. T
- Page 15 and 16: Duchenne muscular dystrophy (DMD).
- Page 17 and 18: WP21 Reachout to the broader RNA co
- Page 19 and 20: 4.1 TABLE OF MILESTONES JPA MONTH 2
- Page 21 and 22: Numerous plasmids, constructs, extr
- Page 23 and 24: collaboration between EURASNET and
- Page 25 and 26: 5.1 WORK PACKAGE REPORTSWork Packag
- Page 27 and 28: • Ongoing regular addition of new
- Page 29 and 30: Work Package 4 The Alternative Spli
- Page 31 and 32: Work Package 6 In silico approaches
- Page 33 and 34: alternative splicing.162 Compare sp
- Page 35 and 36: 33 Agreement on pre-mRNA substrates
- Page 37 and 38: its numerous binding sites on both
- Page 39 and 40: Work Package 8 Genome-wide Analyses
- Page 41 and 42: 173 Further development of data ana
- Page 43 and 44: Problems and explanations for delay
- Page 45 and 46: eveal that there is a dramatic chan
- Page 47 and 48: Work Package 10 Post-translational
- Page 49: mutational analysis that led to the
- Page 53 and 54: FBP11 and 21 with these sequences.
- Page 55 and 56: Similarly to what observed for SF2/
- Page 57 and 58: prp45-113 mutant, which is compatib
- Page 59 and 60: cell lines. In contrast in the pres
- Page 61 and 62: genes with roles in the development
- Page 63 and 64: Structural analyses of several RRMs
- Page 65: Work Package 17 Staff Exchange and
- Page 69 and 70: Barralle's lab, Krämer is acting a
- Page 71 and 72: alternative splicing) and higher ed
- Page 73 and 74: Student Symposium “Decision Makin
- Page 75 and 76: variation and suitability for use w
- Page 77 and 78: Following feedback from members we
- Page 79 and 80: Invited seminars: In addition to me
- Page 81 and 82: Publications 2008Author Title Publi
- Page 83 and 84: Author Title Publication EURASNET P
- Page 85 and 86: Author Title Publication EURASNET P
- Page 87 and 88: Author Title Publication EURASNET P
- Page 89 and 90: Author Title Publication EURASNET P
- Page 91 and 92: Author Title Publication EURASNET P
- Page 93 and 94: Minutes of the General Assembly dec
- Page 96 and 97: 241 Evaluate available careerdevelo
- Page 98 and 99: 156 The EURASNET memberStamm publis
- Page 100 and 101:
y coarse-grained foldingsimulation.
- Page 102 and 103:
overexpressor and mutant lines(mont
- Page 104 and 105:
machinery with particularrespect to
- Page 106 and 107:
alternative splicing, using theEDI
- Page 108 and 109:
contemporary findings withinthe EUR
- Page 110 and 111:
Author Title Publication EURASNET P
- Page 112 and 113:
Author Title Publication EURASNET P
- Page 114 and 115:
Author Title Publication EURASNET P
- Page 116 and 117:
Author Title Publication EURASNET P
- Page 118 and 119:
Licht K, Medenbach J, Lührmann R,K
- Page 120 and 121:
Author Title Publication EURASNET P
- Page 122 and 123:
9 TABULAR OVERVIEW OF MAJOR COST IT
- Page 124 and 125:
total costs * 18.891,12 52778.60 41
- Page 126 and 127:
other (the rest) 48,57 0 56.539,43t
- Page 128 and 129:
travel 6.964,91overhead 3.081,07oth
- Page 130 and 131:
Participant 1A Type of expenditure
- Page 132 and 133:
Participant 1B - Karla Neugebauera)
- Page 134 and 135:
Participant 2 - Juan Valcarcela) Wo
- Page 136 and 137:
Participant 3 - Stefan Stamma) Work
- Page 138 and 139:
Participant 4 - Göran Akusjärvia)
- Page 140 and 141:
Participant 5A/B - Peer Bork/Rolf A
- Page 142 and 143:
Major cost items• ResearchPersonn
- Page 144 and 145:
Participant 7 - Francisco Barallea)
- Page 146 and 147:
Francisco Baralle gave a talk to a
- Page 148 and 149:
alternative splicing on human disea
- Page 150 and 151:
J. Beggs has had frequent contact w
- Page 152 and 153:
Major cost items• ResearchPersonn
- Page 154 and 155:
Major cost items• ResearchPersonn
- Page 156 and 157:
Participant 12A - Jamal Tazia) Work
- Page 158 and 159:
Participant 12B - Bertrand Séraphi
- Page 160 and 161:
Participant 12C - Christiane Branla
- Page 162 and 163:
Participant 12D - Edouard Bertranda
- Page 164 and 165:
Participant 13 - Daniel Schümperli
- Page 166 and 167:
Participant 14 - John W. S. Browna)
- Page 168 and 169:
Participant 15 - Javier F. Cáceres
- Page 170 and 171:
• ResearchPersonnelName WP Person
- Page 172 and 173:
MiscellaneousOligonucleotidesUse of
- Page 174 and 175:
Major cost items• ResearchConsuma
- Page 176 and 177:
Major cost items• ResearchPersonn
- Page 178 and 179:
Participant 21 - Angela Krämera) W
- Page 180 and 181:
Participant 22 - Angus Lamonda) Wor
- Page 182 and 183:
Participant 23 - Chris Smitha) Work
- Page 184 and 185:
ChemicalsEnzymesLaboratory supplies
- Page 186 and 187:
Participant 26 - Didier Auboeufa) W
- Page 188 and 189:
tool for the molecular analysis of
- Page 190 and 191:
Participant 29 - Frédéric Allaina
- Page 192 and 193:
protein and instead possesses the d
- Page 194 and 195:
+ overhead 4.730= total eligible co
- Page 196 and 197:
Claudia Tamaro (PhD) has begun work
- Page 198 and 199:
ETHZIIMCBUPFSOTON-HGDTOTALJoint Pro
- Page 200 and 201:
Direct eligible costs 53.240,48 0,0
- Page 202 and 203:
Direct eligible costs 18.720,07 0,0
- Page 204 and 205:
13 REPORT ON THE DISTRIBUTION OF TH
- Page 206 and 207:
Report on the Distribution of the C
- Page 208 and 209:
For most work packages here follows
- Page 210 and 211:
pathways (eg, Reactome ).While PAND
- Page 212 and 213:
The possible dependence of the effe
- Page 214 and 215:
In some treatments or mutants, simi
- Page 216 and 217:
Work Package 12: Mis-splicing and D
- Page 218 and 219:
acid. Furthermore, using antibodies
- Page 220 and 221:
At the 2007 annual meeting in Ile d
- Page 222 and 223:
Partner 13 has extensively studied
- Page 224 and 225:
inhibitors/modulators of RNA splici
- Page 226 and 227:
18 MONTH JPA: WORKPACKAGE TIMECOURS
- Page 228 and 229:
15.3 GRAPHICAL PRESENTATION OF WORK
- Page 230 and 231:
15.4 TABLE OF MILESTONES (MONTH 37-
- Page 232 and 233:
156 The EURASNET memberStamm publis
- Page 234 and 235:
194 Bertrand will analyze indetail
- Page 236 and 237:
216 Aubeouf and Neugebauer willcoll
- Page 238 and 239:
238 Establishing joint PhDcommittee
- Page 240 and 241:
283 A unified browser for allknown
- Page 242 and 243:
296 Group 12a (Branlant) willuse th
- Page 244 and 245:
315 Lamond will use a FLIM-FRET bas
- Page 246 and 247:
329 GROUP 7 (F. BARALLE)will contin
- Page 248 and 249:
333 GROUP 27 (GABELLINI)will:1) foc
- Page 250 and 251:
345 Schümperli’s group isplannin
- Page 252 and 253:
366 Quarterly network e-mail atmont
- Page 254 and 255:
2. Sharing Resources, Technology an
- Page 256 and 257:
5. Ensuring DurabilityWorkpackage d
- Page 258 and 259:
7. Molecular Characterization of Sp
- Page 260 and 261:
8. Genome-wide Analyses of Splicing
- Page 262 and 263:
9. Complexity of Spliceosomal Prote
- Page 264 and 265:
202 Srebrow will investigate the ro
- Page 266 and 267:
12. Mis-splicing and DiseaseWorkpac
- Page 268 and 269:
13. Co-transcriptional Mechanisms o
- Page 270 and 271:
14. Chemical Biology and Therapeuti
- Page 272 and 273:
15. Development of Enabling Technol
- Page 274 and 275:
269 IFMs in 20091. “Alternative a
- Page 276 and 277:
18. Career DevelopmentWorkpackage d
- Page 278 and 279:
20. SMEs and Technology TransferWor
- Page 280 and 281:
21. Reachout to the Broader RNA Com
- Page 282 and 283:
15.7 WORK PACKAGE LIST (MONTHS 37-5
- Page 284 and 285:
ETHZIIMCBUPFSOTON-HGDTOTALJoint Pro
- Page 286 and 287:
AMU 5 29 8 42UAAR 5 145 16 1665 29
- Page 288 and 289:
Joint Programme of Activities RESEA
- Page 290 and 291:
60 month period 18 month periodRequ
- Page 292 and 293:
Integration1. Young Investigator Pr
- Page 294 and 295:
UCAM-DBIOC 2350,000x1/3 € 50,000x
- Page 296:
the management team. Audit costs ar