Mathematics in Independent Component Analysis
Mathematics in Independent Component Analysis
Mathematics in Independent Component Analysis
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
250 Chapter 18. Proc. EUSIPCO 2006<br />
4.3 Nonnegative Matrix Factorization<br />
(Overcomplete) Nonnegative Matrix Factorization (NMF)<br />
deals with the problem of f<strong>in</strong>d<strong>in</strong>g a nonnegative decomposition<br />
X = AS + N of a nonnegative matrix X, where N<br />
denotes unknown Gaussian noise. S is often pictured as a<br />
source data set conta<strong>in</strong><strong>in</strong>g samples along its columns. If we<br />
assume that S spans the whole first quadrant, then X is a<br />
conic hull with cone l<strong>in</strong>es given by the columns of A. After<br />
projection to the standard simplex, the conic hull reduces<br />
to the convex hull, and the projected, known mixture data<br />
set X lies with<strong>in</strong> a convex polytope of the order given by the<br />
number of rows of S. Hence we face the problem of identify<strong>in</strong>g<br />
edges of a sampled polytope, and, even <strong>in</strong> the overcomplete<br />
case, we may tackle this problem by the Grassmann<br />
cluster<strong>in</strong>g-based identification algorithm from the previous<br />
section.<br />
As an example, see figure 5, we choose a random mix<strong>in</strong>g<br />
matrix<br />
A =<br />
�<br />
0.76 0.39<br />
�<br />
0.14<br />
0.033 0.06 0.43<br />
0.20 0.56 0.43<br />
and sources S given by i.i.d. samples from a squared<br />
gaussian. 10 5 samples were drawn, and 10 to 10 5 sample<br />
subsets where used for the comparison. We refer to the figure<br />
caption for further details.<br />
5. CONCLUSION<br />
We have studied k-means-style cluster<strong>in</strong>g problems on the<br />
non-Euclidean Grassmann manifold. In an adequate metric,<br />
we were able to reduce the aris<strong>in</strong>g centroid calculation<br />
problem to the calculation of eigenvectors of the cluster covariance,<br />
for which we gave a proof based on convex optimization.<br />
The algorithm was illustrated by applications to<br />
polytope fitt<strong>in</strong>g and to perform<strong>in</strong>g overcomplete nonnegative<br />
factorizations similar to NMF. In future work, besides<br />
extend<strong>in</strong>g the framework to other cluster<strong>in</strong>g algorithms and<br />
matrix manifolds together with prov<strong>in</strong>g convergence of the<br />
result<strong>in</strong>g algorithms, we plan on apply<strong>in</strong>g the algorithm for<br />
the stability analysis of multidimensional <strong>in</strong>dependent component<br />
analysis.<br />
Acknowledgements<br />
Partial f<strong>in</strong>ancial support by the DFG (GRK 638) and the<br />
BMBF (project ‘ModKog’) is gratefully acknowledged.<br />
REFERENCES<br />
[1] C.B. Barber, D.P. Dobk<strong>in</strong>, and H. Huhdanpaa. The<br />
quickhull algorithm for convex hull. Technical Report<br />
GCG53, The Geometry Center, University of M<strong>in</strong>nesota,<br />
M<strong>in</strong>neapolis, 1993.<br />
[2] C.M. Bishop. Neural Networks for Pattern Recognition.<br />
Oxford University Press, 1995.<br />
[3] L. Bottou and Y. Bengio. Convergence properties of the<br />
k-means algorithms. In Proc. NIPS 1994, pages 585–<br />
592. MIT Press, 1995.<br />
[4] P.S. Bradley and O.L. Mangasarian. k-plane cluster<strong>in</strong>g.<br />
Journal of Global Optimization, 16(1):23–32, 2000.<br />
Samples<br />
QHull contour<br />
Grasmann cluster<strong>in</strong>g (100 Samples)<br />
NMF Mix<strong>in</strong>g Matrix (100000 Samples)<br />
Mix<strong>in</strong>g matrix<br />
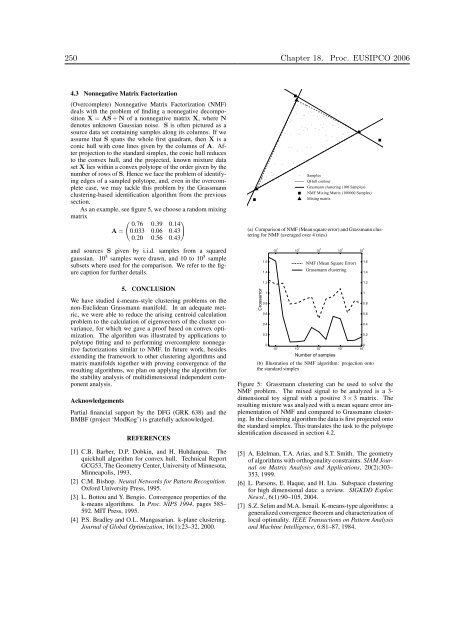
(a) Comparison of NMF (Mean square error) and Grassmann cluster<strong>in</strong>g<br />
for NMF (averaged over 4 tries)<br />
Crosserror<br />
1.6<br />
1.4<br />
1.2<br />
1<br />
0.8<br />
0.6<br />
0.4<br />
0.2<br />
0<br />
10 1<br />
10 1<br />
10 2<br />
10 2<br />
10 3<br />
NMF (Mean Square Error)<br />
Grassmann cluster<strong>in</strong>g<br />
10 3<br />
Number of samples<br />
10 4<br />
10 4<br />
10 5<br />
1.6<br />
1.4<br />
1.2<br />
1<br />
0.8<br />
0.6<br />
0.4<br />
0.2<br />
10 5<br />
0<br />
(b) Illustration of the NMF algorithm: projection onto<br />
the standard simplex<br />
Figure 5: Grassmann cluster<strong>in</strong>g can be used to solve the<br />
NMF problem. The mixed signal to be analyzed is a 3dimensional<br />
toy signal with a positive 3 × 3 matrix. The<br />
result<strong>in</strong>g mixture was analyzed with a mean square error implementation<br />
of NMF and compared to Grassmann cluster<strong>in</strong>g.<br />
In the cluster<strong>in</strong>g algorithm the data is first projected onto<br />
the standard simplex. This translates the task to the polytope<br />
identification discussed <strong>in</strong> section 4.2.<br />
[5] A. Edelman, T.A. Arias, and S.T. Smith. The geometry<br />
of algorithms with orthogonality constra<strong>in</strong>ts. SIAM Journal<br />
on Matrix <strong>Analysis</strong> and Applications, 20(2):303–<br />
353, 1999.<br />
[6] L. Parsons, E. Haque, and H. Liu. Subspace cluster<strong>in</strong>g<br />
for high dimensional data: a review. SIGKDD Explor.<br />
Newsl., 6(1):90–105, 2004.<br />
[7] S.Z. Selim and M.A. Ismail. K-means-type algorithms: a<br />
generalized convergence theorem and characterization of<br />
local optimality. IEEE Transactions on Pattern <strong>Analysis</strong><br />
and Mach<strong>in</strong>e Intelligence, 6:81–87, 1984.