Damm et al.combined sequence datasets. Models of nucleotide substitutionf<strong>or</strong> each gene determined by MrModeltest v. 2.3 were included f<strong>or</strong>each gene partition. The analyses of two MCMC chains were runfrom random trees f<strong>or</strong> one million generations and sampled every100 generations. The likelihood sc<strong>or</strong>e of the two runs were 2 730and 2 170 and theref<strong>or</strong>e, the first 2 450 (the average of both) treeswere discarded as the burn-in phase of the analysis and posteri<strong>or</strong>probabilities determined from the remaining trees. Sequencesderived in this study were lodged at GenBank, the alignmentin TreeBASE (www.treebase.<strong>or</strong>g), and taxonomic novelties inMycoBank (Crous et al. 2004a).RESULTSPhylogenyThe seven sequence datasets did not show any conflicts in treetopology f<strong>or</strong> the 70 % reciprocal bootstrap trees, which allowed usto combine them. In the multigene analyses (gene boundaries ofITS: 1–561, GAPDH: 572–864, CHS-1: 875–1154, HIS3: 1164–1562, ACT: 1573–1851, TUB2: 1862–2363, CAL: 2374–2823) of87 isolates of C. boninense and related <strong>Colletotrichum</strong> <strong>species</strong>including the outgroup, 2 823 characters including the alignmentgaps were processed, of which 572 characters were parsimonyinf<strong>or</strong>mative,247 parsimony-uninf<strong>or</strong>mative and 2004 constant. Aftera heuristic search using PAUP, 958 most parsimonious trees wereretained (length = 1423 steps, CI = 0.740, RI = 0.927, RC = 0.686,HI = 0.260) of which one is shown in Fig. 1. The topology of the 958trees was similar, which was verified f<strong>or</strong> a large selection of trees.They differed in the position of taxa within the subclades and in theposition of strain <strong>CBS</strong> 130241. F<strong>or</strong> Bayesian analysis, a HKY+Gmodel was selected f<strong>or</strong> ACT and CAL, SYM+I+G f<strong>or</strong> ITS, K80+I+Gf<strong>or</strong> CHS-1, GTR+I+G f<strong>or</strong> HIS3, and HKY+I f<strong>or</strong> GAPDH and TUB2,and inc<strong>or</strong>p<strong>or</strong>ated in the analysis. The consensus tree obtainedfrom Bayesian analyses confirmed the tree topology obtained withparsimony as well as the bootstrap supp<strong>or</strong>t (Fig. 1).The analyses resulted in the detection of 18 clades, whichwe accept as representing different <strong>Colletotrichum</strong> <strong>species</strong>.M<strong>or</strong>e than half of all strains included cluster in the first clade (C.karstii) with a bootstrap supp<strong>or</strong>t of 96 % and a Bayesian posteri<strong>or</strong>probability value of 1.00. Two single strain clades, representingC. phyllanthi and C. annellatum, group with this big clade with abootstrap supp<strong>or</strong>t/Bayesian posteri<strong>or</strong> probability value of 96/0.99and 100/1.00, respectively. The C. petchii clade is well supp<strong>or</strong>ted(100/1.00) and f<strong>or</strong>ms a sister clade to the first three <strong>species</strong>. Theclade representing C. novae-zelandiae consists of two strains ona long branch (100/1.00). In contrast, the following five clades aresh<strong>or</strong>t-branched, namely C. boninense s. str. (95/1.00), C. t<strong>or</strong>ulosum(100/1.00), C. cymbidiicola (96/1.00), C. oncidii (100/1.00) and aclade containing an unnamed single strain (<strong>CBS</strong> 123921). Thesefive <strong>species</strong> f<strong>or</strong>m a sister clade (100/1.00) to another well supp<strong>or</strong>ted(100/1.00) clade f<strong>or</strong>med by two single strain clades, C. beeveriand C. brassicicola, and the C. colombiense clade (100/1.00)containing two strains. The clades representing C. hippeastri andC. brasiliense consist of three and two strains respectively, andare well supp<strong>or</strong>ted (100/1.00). They group (100/1.00) with a singlestrain clade (C. parsonsiae). The C. constrictum clade (100/1.00)containing two strains and the single strain clade representing C.dacrycarpi are on very long branches and group with a Bayesianposteri<strong>or</strong> probability value of 1.00.The individual alignments and trees of the seven single geneswere compared as well with respect to their perf<strong>or</strong>mance in <strong>species</strong>recognition. With ITS and CHS-1 only 7 and 9 <strong>species</strong> can berecognised, with TUB2 some <strong>species</strong> close to C. boninense cannot be separated, while with HIS3 and CAL some <strong>species</strong> onlydiffer in one <strong>or</strong> two bp and f<strong>or</strong>m no <strong>or</strong> only sh<strong>or</strong>t-branched clades.With GAPDH all clades are recognised, but some are also sh<strong>or</strong>tbranched,especially the single strain clades C. beeveri and C.brassicicola that are well differentiated with almost all other genesexcept f<strong>or</strong> ITS. With ACT the intraspecific variability is very high insome <strong>species</strong>, which could lead to misidentifications.TaxonomyThe 86 isolates studied (Table 1) are assigned to 18 <strong>species</strong> withinthe <strong>Colletotrichum</strong> boninense <strong>complex</strong> based on DNA sequencedata and m<strong>or</strong>phology, including 12 <strong>species</strong> that are new to science.Ten <strong>species</strong> f<strong>or</strong>m teleom<strong>or</strong>ph stages in vitro, four <strong>species</strong> haveknown anam<strong>or</strong>phs described in <strong>Colletotrichum</strong>, while one <strong>species</strong>,G. phyllanthi, has a known teleom<strong>or</strong>ph and is shown here asbelonging to the <strong>Colletotrichum</strong> boninense <strong>species</strong> <strong>complex</strong>. All<strong>species</strong> studied in culture are characterised below.Species of <strong>Colletotrichum</strong> represent anam<strong>or</strong>phic stages ofGlomerella. Anam<strong>or</strong>ph and telem<strong>or</strong>ph names of fungi will haveequal status under the International Code of Nomenclature f<strong>or</strong>algae, fungi, and plants (f<strong>or</strong>merly the International Code f<strong>or</strong>Botanical Nomenclature), with the deletion of Art. 59, which takeseffect on 1 Jan. 2013. The decision was qualified by a stipulationthat uptake of names of anam<strong>or</strong>phic genera that predate well-knowncompeting teleom<strong>or</strong>phic names should be ratified by a committeeof the International Commission f<strong>or</strong> the Taxonomy of Fungi. Thename <strong>Colletotrichum</strong> (C<strong>or</strong>da 1831) predates Glomerella (Spauld.& H. Schrenk 1903) and is m<strong>or</strong>e commonly used. Consequentlywe name the new holom<strong>or</strong>phs as <strong>species</strong> of <strong>Colletotrichum</strong> and wedo not name the new sexual m<strong>or</strong>phs separately. Furtherm<strong>or</strong>e, G.phyllanthi is combined in <strong>Colletotrichum</strong> as C. phyllanthi. There isprecedent f<strong>or</strong> this as Rojas et al. (2010) described <strong>Colletotrichum</strong>ignotum as having a teleom<strong>or</strong>ph (see Rojas et al. 2010: figs 44–47,table III), although the teleom<strong>or</strong>phic structures were not included inthe f<strong>or</strong>mal <strong>species</strong> diagnosis.<strong>Colletotrichum</strong> annellatum Damm, P.F. Cannon & Crous,sp. nov. MycoBank MB560734. Fig. 2.Etymology: The name refers to the proliferation of conidiogenouscells, which appear annellate.Teleom<strong>or</strong>ph developed on SNA. Ascomata ovoid to obpyrif<strong>or</strong>m,medium to dark brown, 180–220 × 100–150 µm, glabrous,ostiolate, neck hyaline to pale brown, wall 5–10 µm thick, outerlayer composed of flattened medium brown angular cells, 5–10 µmdiam. Interascal tissue composed of paraphyses, hyaline, septate,branched at the base, disintegrating quickly, 35–55 µm long, base3–4.5 µm diam, apically free, the apex rounded. Asci cylindricalto clavate, 58–74 × 11–16 µm, 8-sp<strong>or</strong>ed. Ascosp<strong>or</strong>es arrangedbiseriately, hyaline, smooth-walled, aseptate, cylindrical to narrowlyfabif<strong>or</strong>m, straight <strong>or</strong> rarely very slightly curved, both sides rounded,(13.5–)15–17(–18.5) × 5–6 µm, mean ± SD = 16.0 ± 1.1 × 5.6 ±0.4 µm, L/W ratio = 2.9.Teleom<strong>or</strong>ph developed on Anthriscus stem. Ascomata ovoidto obpyrif<strong>or</strong>m, medium brown. Asci cylindrical to clavate, 60–706
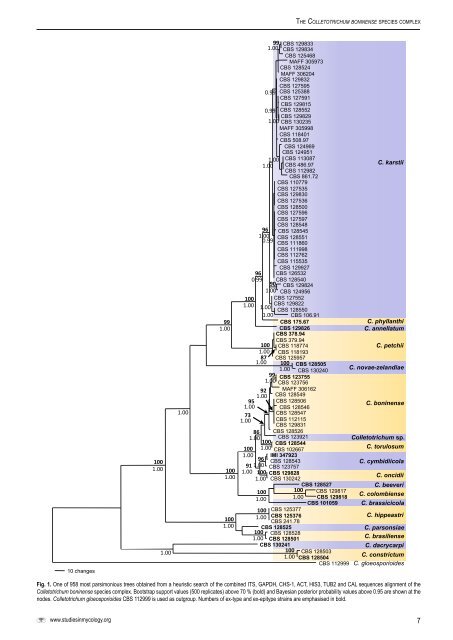
The <strong>Colletotrichum</strong> boninense <strong>species</strong> <strong>complex</strong>10 changes1001.001.001.00991.001001.001001.00921.00951.00731.0099 <strong>CBS</strong> 1298331.00 <strong>CBS</strong> 129834<strong>CBS</strong> 125468MAFF 305973<strong>CBS</strong> 128524MAFF 306204<strong>CBS</strong> 129832<strong>CBS</strong> 1275950.99 <strong>CBS</strong> 125388<strong>CBS</strong> 127591<strong>CBS</strong> 1298150.99 <strong>CBS</strong> 128552<strong>CBS</strong> 1298291.00 <strong>CBS</strong> 130235MAFF 305998<strong>CBS</strong> 118401<strong>CBS</strong> 508.97<strong>CBS</strong> 124969<strong>CBS</strong> 1249511.00 <strong>CBS</strong> 1130871.00 <strong>CBS</strong> 486.97<strong>CBS</strong> 112982<strong>CBS</strong> 861.72<strong>CBS</strong> 110779<strong>CBS</strong> 127535<strong>CBS</strong> 129830<strong>CBS</strong> 127536<strong>CBS</strong> 128500<strong>CBS</strong> 127596<strong>CBS</strong> 127597<strong>CBS</strong> 12854896 <strong>CBS</strong> 1285451.00 <strong>CBS</strong> 1285510.99 <strong>CBS</strong> 111860<strong>CBS</strong> 111998<strong>CBS</strong> 112762<strong>CBS</strong> 115535<strong>CBS</strong> 12992796 <strong>CBS</strong> 1265320.99 <strong>CBS</strong> 12854090 <strong>CBS</strong> 1298241.00 <strong>CBS</strong> 124956100 <strong>CBS</strong> 1275521.00 <strong>CBS</strong> 1298221.001.00<strong>CBS</strong> 128550<strong>CBS</strong> 106.91<strong>CBS</strong> 175.67<strong>CBS</strong> 129826<strong>CBS</strong> 378.94<strong>CBS</strong> 379.94<strong>CBS</strong> 118774<strong>CBS</strong> 1181931001.0087 <strong>CBS</strong> 1259571.00 100 <strong>CBS</strong> 1285051.00 <strong>CBS</strong> 13024099 <strong>CBS</strong> 1237551.00 <strong>CBS</strong> 123756MAFF 306162<strong>CBS</strong> 128549<strong>CBS</strong> 128506<strong>CBS</strong> 128546<strong>CBS</strong> 128547<strong>CBS</strong> 112115<strong>CBS</strong> 129831<strong>CBS</strong> 128526<strong>CBS</strong> 123921C. karstiiC. phyllanthiC. annellatumC. petchiiC. novae-zelandiaeC. boninense861.00<strong>Colletotrichum</strong> sp.100 <strong>CBS</strong> 128544100 1.00C. t<strong>or</strong>ulosum<strong>CBS</strong> 1026671.00 IMI 34792396 <strong>CBS</strong> 128543C. cymbidiicola91 1.00 <strong>CBS</strong> 1237571.00 100 <strong>CBS</strong> 129828C. oncidii1.00 <strong>CBS</strong> 130242<strong>CBS</strong> 128527C. beeveri100 100 <strong>CBS</strong> 1298171.00 1.00C. colombiense<strong>CBS</strong> 129818<strong>CBS</strong> 101059 C. brassicicola100 <strong>CBS</strong> 1253771.00 <strong>CBS</strong> 125376C. hippeastri<strong>CBS</strong> 241.78<strong>CBS</strong> 128525C. parsonsiae100 <strong>CBS</strong> 1285281.00 <strong>CBS</strong> 128501C. brasiliense<strong>CBS</strong> 130241C. dacrycarpi100 <strong>CBS</strong> 1285031.00C. constrictum<strong>CBS</strong> 128504<strong>CBS</strong> 112999 C. gloeosp<strong>or</strong>ioidesFig. 1. One of 958 most parsimonious trees obtained from a heuristic search of the combined ITS, GAPDH, CHS-1, ACT, HIS3, TUB2 and CAL sequences alignment of the<strong>Colletotrichum</strong> boninense <strong>species</strong> <strong>complex</strong>. Bootstrap supp<strong>or</strong>t values (500 replicates) above 70 % (bold) and Bayesian posteri<strong>or</strong> probability values above 0.95 are shown at thenodes. <strong>Colletotrichum</strong> gloeosp<strong>or</strong>ioides <strong>CBS</strong> 112999 is used as outgroup. Numbers of ex-type and ex-epitype strains are emphasised in bold.www.studiesinmycology.<strong>or</strong>g7
- Page 1: Studies in Mycology 73 (September 2
- Page 5 and 6: Colletotrichum: complex species or
- Page 10 and 11: Damm et al.Table 1. Strains of Coll
- Page 12 and 13: Damm et al.Table 1. (Continued).Gen
- Page 16 and 17: Damm et al.Fig. 2. Colletotrichum a
- Page 19 and 20: The Colletotrichum boninense specie
- Page 21 and 22: The Colletotrichum boninense specie
- Page 23 and 24: The Colletotrichum boninense specie
- Page 25 and 26: The Colletotrichum boninense specie
- Page 27 and 28: The Colletotrichum boninense specie
- Page 29 and 30: The Colletotrichum boninense specie
- Page 31 and 32: The Colletotrichum boninense specie
- Page 33 and 34: The Colletotrichum boninense specie
- Page 35 and 36: The Colletotrichum boninense specie
- Page 37 and 38: The Colletotrichum boninense specie
- Page 39 and 40: The Colletotrichum boninense specie
- Page 41 and 42: The Colletotrichum boninense specie
- Page 43 and 44: The Colletotrichum boninense specie
- Page 45 and 46: available online at www.studiesinmy
- Page 47 and 48: The Colletotrichum acutatum species
- Page 49 and 50: The Colletotrichum acutatum species
- Page 51 and 52: The Colletotrichum acutatum species
- Page 53 and 54: The Colletotrichum acutatum species
- Page 55 and 56: The Colletotrichum acutatum species
- Page 57 and 58: The Colletotrichum acutatum species
- Page 59 and 60: The Colletotrichum acutatum species
- Page 61 and 62: The Colletotrichum acutatum species
- Page 63 and 64: The Colletotrichum acutatum species
- Page 65 and 66:
The Colletotrichum acutatum species
- Page 67 and 68:
The Colletotrichum acutatum species
- Page 69 and 70:
The Colletotrichum acutatum species
- Page 71 and 72:
The Colletotrichum acutatum species
- Page 73 and 74:
The Colletotrichum acutatum species
- Page 75 and 76:
The Colletotrichum acutatum species
- Page 77 and 78:
The Colletotrichum acutatum species
- Page 79 and 80:
The Colletotrichum acutatum species
- Page 81 and 82:
The Colletotrichum acutatum species
- Page 83 and 84:
The Colletotrichum acutatum species
- Page 85 and 86:
The Colletotrichum acutatum species
- Page 87 and 88:
The Colletotrichum acutatum species
- Page 89 and 90:
The Colletotrichum acutatum species
- Page 91:
The Colletotrichum acutatum species
- Page 94 and 95:
Damm et al.µm diam. Setae not obse
- Page 96 and 97:
Damm et al.Table 2. Conidia measure
- Page 98 and 99:
Damm et al.Table 2. (Continued).Spe
- Page 100 and 101:
Damm et al.Table 3. Appressoria mea
- Page 102 and 103:
Damm et al.Fig. 25. Colletotrichum
- Page 104 and 105:
Damm et al.Fig. 26. Colletotrichum
- Page 106 and 107:
Damm et al.Fig. 27. Colletotrichum
- Page 108 and 109:
Damm et al.Fig. 28. Colletotrichum
- Page 110 and 111:
Damm et al.Fig. 29. Colletotrichum
- Page 112 and 113:
Damm et al.Fig. 30. Colletotrichum
- Page 114 and 115:
Damm et al.to olivaceous grey, filt
- Page 116 and 117:
Damm et al.are formed directly from
- Page 118 and 119:
Damm et al.Arx JA von (1957). Die A
- Page 120 and 121:
Damm et al.Masaba DM, Waller JM (19
- Page 122 and 123:
114
- Page 124 and 125:
Weir et al.grass-inhabiting species
- Page 126 and 127:
Weir et al.Table 1. A list of strai
- Page 128 and 129:
Weir et al.Table 1. (Continued).Spe
- Page 130 and 131:
Weir et al.Table 1. (Continued).Spe
- Page 132 and 133:
Weir et al.Table 2. Primers used in
- Page 134 and 135:
Weir et al.11ICMP 17789 Malus USA1I
- Page 136 and 137:
Weir et al.Kahawae cladeC. alataeC.
- Page 138 and 139:
Weir et al.A0.991ICMP 17905 Coffea
- Page 140 and 141:
Weir et al.Conidial width variation
- Page 142 and 143:
Weir et al.G. c.“f.sp. camelliae
- Page 144 and 145:
Weir et al.Fig. 10. Colletotrichum
- Page 146 and 147:
Weir et al.Fig. 12. Colletotrichum
- Page 148 and 149:
Weir et al.Fig. 13. Colletotrichum
- Page 150 and 151:
Weir et al.Fig. 15. Colletotrichum
- Page 152 and 153:
Weir et al.Fig. 17. Colletotrichum
- Page 154 and 155:
Weir et al.Fig. 18. Glomerella cing
- Page 156 and 157:
Weir et al.images under C. fructico
- Page 158 and 159:
Weir et al.Specimen examined: Thail
- Page 160 and 161:
Weir et al.Fig. 23. Colletotrichum
- Page 162 and 163:
Weir et al.Notes: First described f
- Page 164 and 165:
Weir et al.gossypii var. cephalospo
- Page 166 and 167:
Weir et al.to utilise either citrat
- Page 168 and 169:
Weir et al.Fig. 27. Colletotrichum
- Page 170 and 171:
Weir et al.(ex-epitype culture CBS
- Page 172 and 173:
Weir et al.of this species to C. ps
- Page 174 and 175:
Weir et al.Fig. 32. Colletotrichum
- Page 176 and 177:
Weir et al.Fig. 33. Colletotrichum
- Page 178 and 179:
Weir et al.Fig. 35. Colletotrichum
- Page 180 and 181:
Weir et al.Fig. 36. Colletotrichum
- Page 182 and 183:
Weir et al.Fig. 38. Colletotrichum
- Page 184 and 185:
Weir et al.Table 4. Performance of
- Page 186 and 187:
Weir et al.Crous PW, Gams W, Stalpe
- Page 188 and 189:
Weir et al.Simmonds JH (1968). Type
- Page 190 and 191:
Cannon et al.182
- Page 192 and 193:
Cannon et al.In rare instances, Col
- Page 194 and 195:
Cannon et al.Table 1. Summary of pr
- Page 196 and 197:
Cannon et al.Table 1. (Continued).P
- Page 198 and 199:
Cannon et al.Table 2. Colletotrichu
- Page 200 and 201:
Cannon et al.as did those for Colle
- Page 202 and 203:
Cannon et al.Table 3. Authentic seq
- Page 204 and 205:
Cannon et al.Table 3. (Continued).S
- Page 206 and 207:
Cannon et al.Table 3. (Continued).S
- Page 208 and 209:
Cannon et al.Table 3. (Continued).S
- Page 210 and 211:
Cannon et al.110.74 C. cuscutae IMI
- Page 212 and 213:
Cannon et al.Boninense cladeThe bon
- Page 214 and 215:
Cannon et al.C. lindemuthianum AB08
- Page 216 and 217:
Cannon et al.A further important ac
- Page 218 and 219:
Cannon et al.Hawksworth DL (2011).
- Page 220 and 221:
Cannon et al.Rodríguez-Guerra R, R
- Page 223 and 224:
Studies in MycologyStudies in Mycol
- Page 225 and 226:
CBS Biodiversity SeriesNo. 11:Taxon
- Page 227:
Studies in Mycology (ISSN 0166-0616