Weir et al.gossypii var. cephalosp<strong>or</strong>ioides are ITS2 and the D2 region ofthe rDNA LSU, neither of which resolves their relationships withinthe C. gloeosp<strong>or</strong>ioides <strong>complex</strong>. Whether the seedling pathogenregarded by Silva-Mann et al. (2005) and Bailey et al. (1996) to beC. gossypii represents the <strong>species</strong> first described from cotton inthe USA is not known. The genetic relationship of these apparentlybiologically specialised fungi requires additional sequences to begenerated from authentic isolates with known pathogenicity.<strong>Colletotrichum</strong> gossypii var. cephalosp<strong>or</strong>ioides A.S.Costa, Bragantia 6: 5. 1946.≡ <strong>Colletotrichum</strong> gloeosp<strong>or</strong>ioides “var. cephalosp<strong>or</strong>ioides” (A.S. Costa)Follin & Mangano, Coton et fibres tropicales 37: 209. 1983. [comb. inval.,no full reference to basionym]Notes: See notes under <strong>Colletotrichum</strong> gossypii.* <strong>Colletotrichum</strong> h<strong>or</strong>ii B. Weir & P.R. Johnst., Mycotaxon111: 21. 2010.Weir & Johnston (2010) and Xie et al. (2010a) provide descriptions.Geographic distribution and host range: Associated with fruit andstem disease of Diospyros kaki from China, Japan, and NewZealand. Xie et al. (2010a) noted min<strong>or</strong> symptoms on inoculatedfruit of Capsicum annuum, Musa acuminata, and Cucurbita pepo,but noted that the fungus had never been associated with diseasesymptoms on these hosts from the field.Genetic identification: ITS distinguishes C. h<strong>or</strong>ii from all other<strong>species</strong>.Specimens examined: See Weir & Johnston (2010).<strong>Colletotrichum</strong> hymenocallidis Yan L. Yang, Zuo Y. Liu,K.D. Hyde & L. Cai, Fungal Diversity 39: 138. 2009.Notes: Placed here in synonymy with <strong>Colletotrichum</strong> siamense.See notes and additional specimens examined under C. siamense.Yang et al. (2009) rep<strong>or</strong>ted this <strong>species</strong> as a leaf pathogen ofHymenocallis americana. They distinguished C. hymenocallidisfrom C. siamense, also described from Hymenocallis, primarilyon the basis of a multi-gene phylogeny and differences in colonycolour. Although gene selection was appropriate f<strong>or</strong> resolvinggenetic relationships within the C. gloeosp<strong>or</strong>ioides group, Yanget al. (2009) included only five isolates of the C. gloeosp<strong>or</strong>ioides<strong>complex</strong> in their phylogeny. Based on this isolate selection, theC. hymenocallidis isolates were genetically distinct from theC. siamense isolates. However, in our analysis, in which the C.siamense/C. hymenocallidis group is represented by 30 isolatesfrom a wide range of hosts from all over the w<strong>or</strong>ld, authentic isolatesof the two <strong>species</strong> fall within a monophyletic clade that cannot befurther subdivided phylogenetically.The Latin part of the C. hymenocallidis protologue designatesa culture (“Holotypus: Cultura (CSSN2)”) as the holotype butthe English citation of the type specimen c<strong>or</strong>rects this apparentmistake, citing CSSN2 as an ex-holotype culture, with the herbariumspecimen GZAAS 080001 as the holotype.Specimen examined: China, Guangxi, Nanning, on Hymenocallis americana leafspot, coll. Y.L. Yang GZAAS 080001, 19 Jun 2008 (ex-holotype culture of C.hymenocallidis − <strong>CBS</strong> 125378 = ICMP 18642).<strong>Colletotrichum</strong> ignotum E.I Rojas, S.A. Rehner & Samuels,Mycologia 102: 1331. 2010.Notes: Placed here in synonymy with <strong>Colletotrichum</strong> fructicola. Seenotes and additional specimens examined under C. fructicola.Specimen examined: Panama: Barro Col<strong>or</strong>ado Monument, Tetragastris panamensisleaf endophyte, coll. E.I. Rojas E886, 1 Jun 2004 (ex-holotype culture of C.ignotum − <strong>CBS</strong> 125397 = ICMP 18646).<strong>Colletotrichum</strong> jasmini-sambac Wikee, K.D. Hyde, L. Cai& McKenzie, Fungal Diversity 46: 174. 2011.Notes: Placed here in synonymy with <strong>Colletotrichum</strong> siamensebased on the ITS, GAPDH, CAL, TUB2, and ACT gene sequencesfrom the ex-holotype culture, deposited in GenBank by Wikee etal. (2011).Wikee et al. (2011) discussed similarities between C. jasminisambac,C. siamense and C. hymenocallidis, three <strong>species</strong>genetically close in their phylogenetic analysis. The broader rangeof isolates representing C. siamense in our analysis shows thatthese <strong>species</strong> f<strong>or</strong>m a single, monophyletic clade that cannot besensibly subdivided (see notes under C. siamense).Specimen examined: Vietnam, Cu Chi District, Trung An Ward, on living leaves ofJasminum sambac, Jan. 2009, coll. Hoa Nguyen Thi LLTA–01 (ex-holotype cultureof C. jasmini-sambac – <strong>CBS</strong> 130420 = ICMP 19118).* <strong>Colletotrichum</strong> kahawae J.M. Waller & Bridge subsp.kahawae, Mycol. Res. 97: 993. 1993. Fig. 25.Waller et al. (1993) provide a description.Geographic distribution and host range: Known only from Coffeafrom Africa.Genetic identification: ACT, CAL, CHS-1, GAPDH, TUB2, SOD2,and ITS sequences are the same as those from C. kahawaesubsp. ciggaro. The two sub<strong>species</strong> can be distinguished by GSsequences; C. kahawae subsp. kahawae has a 22 bp deletion anda single C to T transition. Collectively, the two sub<strong>species</strong> can bedistinguished from all other <strong>species</strong> using ITS sequences alone.Notes: <strong>Colletotrichum</strong> kahawae was proposed by Waller et al. (1993)as a name to refer specifically to <strong>Colletotrichum</strong> isolates causingCoffee Berry Disease (CBD), to taxonomically distinguish thesedisease-causing isolates from the several other <strong>Colletotrichum</strong>spp. that can be isolated from coffee plants, including C. coffeanum(see notes under C. coffeanum). <strong>Colletotrichum</strong> kahawae is anapparently clonal population (Varzea et al. 2002), widespread oncoffee in Africa, and with a distinctive growth f<strong>or</strong>m and biology(Waller et al. 1993).In this paper C. kahawae sensu Waller et al. (1993) is reducedto sub<strong>species</strong>. Based on ACT, CAL, CHS-1, GAPDH, TUB2, SOD2,and ITS gene sequences the coffee berry pathogen cannot bedistinguished from isolates from a wide range of other hosts thatare not pathogenic to coffee. Those other isolates are referred tohere as C. kahawae subsp. ciggaro. We retain a distinct taxonomiclabel f<strong>or</strong> the coffee berry pathogen to reflect its biosecurityimp<strong>or</strong>tance. In addition to its biology, C. kahawae subsp. kahawaecan be distinguished metabolically, and genetically using GS genesequences. Waller et al. (1993) used a metabolic test, an inability156
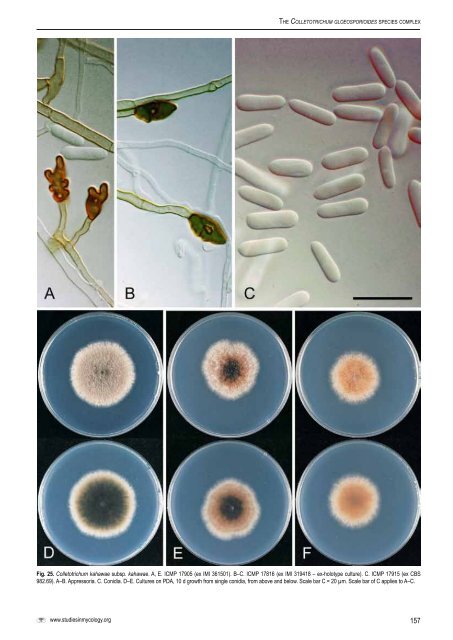
The <strong>Colletotrichum</strong> gloeosp<strong>or</strong>ioides <strong>species</strong> <strong>complex</strong>Fig. 25. <strong>Colletotrichum</strong> kahawae subsp. kahawae. A, E. ICMP 17905 (ex IMI 361501). B–C. ICMP 17816 (ex IMI 319418 – ex-holotype culture). C. ICMP 17915 (ex <strong>CBS</strong>982.69). A–B. Appress<strong>or</strong>ia. C. Conidia. D–E. Cultures on PDA, 10 d growth from single conidia, from above and below. Scale bar C = 20 µm. Scale bar of C applies to A–C.www.studiesinmycology.<strong>or</strong>g157