Windows QTL Cartographer 2.5 - FTP Directory Listing
Windows QTL Cartographer 2.5 - FTP Directory Listing
Windows QTL Cartographer 2.5 - FTP Directory Listing
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
64<br />
<strong>Windows</strong> <strong>QTL</strong> <strong>Cartographer</strong> <strong>2.5</strong><br />
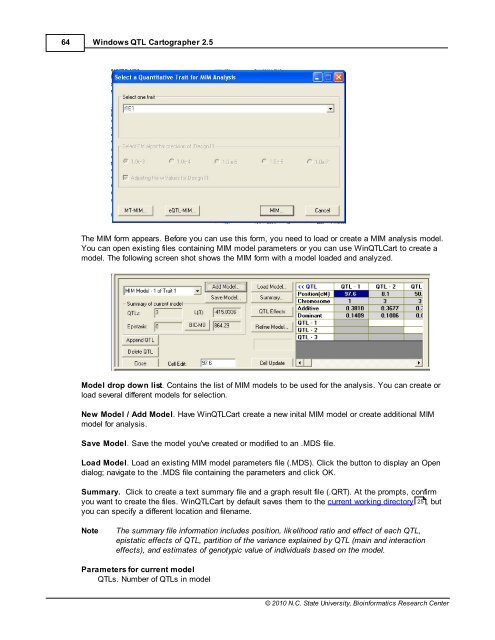
The MIM form appears. Before you can use this form, you need to load or create a MIM analysis model.<br />
You can open existing files containing MIM model parameters or you can use Win<strong>QTL</strong>Cart to create a<br />
model. The following screen shot shows the MIM form with a model loaded and analyzed.<br />
Model drop down list. Contains the list of MIM models to be used for the analysis. You can create or<br />
load several different models for selection.<br />
New Model / Add Model. Have Win<strong>QTL</strong>Cart create a new inital MIM model or create additional MIM<br />
model for analysis.<br />
Save Model. Save the model you've created or modified to an .MDS file.<br />
Load Model. Load an existing MIM model parameters file (.MDS). Click the button to display an Open<br />
dialog; navigate to the .MDS file containing the parameters and click OK.<br />
Summary. Click to create a text summary file and a graph result file (.QRT). At the prompts, confirm<br />
you want to create the files. Win<strong>QTL</strong>Cart by default saves them to the current working directory 25<br />
, but<br />
you can specify a different location and filename.<br />
Note The summary file information includes position, likelihood ratio and effect of each <strong>QTL</strong>,<br />
epistatic effects of <strong>QTL</strong>, partition of the variance explained by <strong>QTL</strong> (main and interaction<br />
effects), and estimates of genotypic value of individuals based on the model.<br />
Parameters for current model<br />
<strong>QTL</strong>s. Number of <strong>QTL</strong>s in model<br />
© 2010 N.C. State University, Bioinformatics Research Center