Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
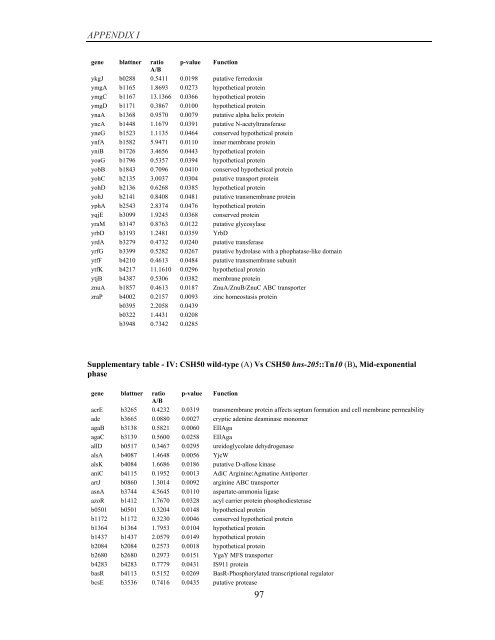
APPENDIX I<br />
<strong>gene</strong> blattner ratio p-value Function<br />
A/B<br />
ykgJ b0288 0.5411 0.0198 putative ferredoxin<br />
ymgA b1165 1.8693 0.0273 hypothetical protein<br />
ymgC b1167 13.1366 0.0366 hypothetical protein<br />
ymgD b1171 0.3867 0.0100 hypothetical protein<br />
ynaA b1368 0.9570 0.0079 putative alpha helix protein<br />
yncA b1448 1.1679 0.0391 putative N-acetyltransferase<br />
yneG b1523 1.1135 0.0464 conserved hypothetical protein<br />
ynfA b1582 5.9471 0.0110 inner membrane protein<br />
yniB b1726 3.4656 0.0443 hypothetical protein<br />
yoaG b1796 0.5357 0.0394 hypothetical protein<br />
yobB b1843 0.7096 0.0410 conserved hypothetical protein<br />
yohC b2135 3.0037 0.0304 putative transport protein<br />
yohD b2136 0.6268 0.0385 hypothetical protein<br />
yohJ b2141 0.8408 0.0481 putative transmembrane protein<br />
yphA b2543 2.8374 0.0476 hypothetical protein<br />
yqjE b3099 1.9245 0.0368 conserved protein<br />
yraM b3147 0.8763 0.0122 putative glycosylase<br />
yrbD b3193 1.2481 0.0359 YrbD<br />
yrdA b3279 0.4732 0.0240 putative transferase<br />
yrfG b3399 0.5282 0.0267 putative hydrolase with a phophatase-like domain<br />
ytfF b4210 0.4613 0.0484 putative transmembrane subunit<br />
ytfK b4217 11.1610 0.0296 hypothetical protein<br />
ytjB b4387 0.5306 0.0382 membrane protein<br />
znuA b1857 0.4613 0.0187 ZnuA/ZnuB/ZnuC ABC transporter<br />
zraP b4002 0.2157 0.0093 zinc homeostasis protein<br />
b0395 2.2058 0.0439<br />
b0322 1.4431 0.0208<br />
b3948 0.7342 0.0285<br />
Supplementary table - IV: CSH50 wild-type (A) Vs CSH50 hns-205::Tn10 (B), Mid-exponential<br />
phase<br />
<strong>gene</strong> blattner ratio p-value Function<br />
A/B<br />
acrE b3265 0.4232 0.0319 transmembrane protein affects septum formation and cell membrane permeability<br />
ade b3665 0.0880 0.0027 cryptic adenine deaminase monomer<br />
agaB b3138 0.5821 0.0060 EIIAga<br />
agaC b3139 0.5600 0.0258 EIIAga<br />
allD b0517 0.3467 0.0295 ureidoglycolate dehydrogenase<br />
alsA b4087 1.4648 0.0056 YjcW<br />
alsK b4084 1.6686 0.0186 putative D-allose kinase<br />
aniC b4115 0.1952 0.0013 AdiC Arginine:Agmatine Antiporter<br />
artJ b0860 1.3014 0.0092 arginine ABC transporter<br />
asnA b3744 4.5645 0.0110 aspartate-ammonia ligase<br />
azoR b1412 1.7670 0.0328 acyl carrier protein phosphodiesterase<br />
b0501 b0501 0.3204 0.0148 hypothetical protein<br />
b1172 b1172 0.3230 0.0046 conserved hypothetical protein<br />
b1364 b1364 1.7953 0.0104 hypothetical protein<br />
b1437 b1437 2.0579 0.0149 hypothetical protein<br />
b2084 b2084 0.2573 0.0018 hypothetical protein<br />
b2680 b2680 0.2973 0.0151 YgaY MFS transporter<br />
b4283 b4283 0.7779 0.0431 IS911 protein<br />
basR b4113 0.5152 0.0269 BasR-Phosphorylated transcriptional regulator<br />
bcsE b3536 0.7416 0.0435 putative protease<br />
97