Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
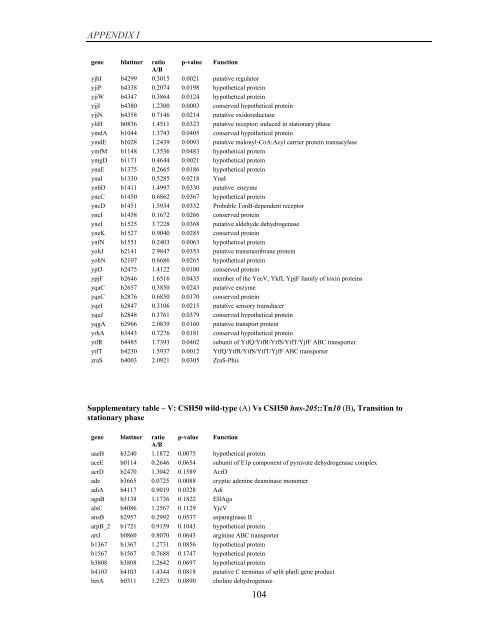
APPENDIX I<br />
<strong>gene</strong> blattner ratio p-value Function<br />
A/B<br />
yjhI b4299 0.3015 0.0021 putative regulator<br />
yjiP b4338 0.2074 0.0198 hypothetical protein<br />
yjiW b4347 0.3864 0.0124 hypothetical protein<br />
yjjI b4380 1.2300 0.0003 conserved hypothetical protein<br />
yjjN b4358 0.7146 0.0214 putative oxidoreductase<br />
yliH b0836 1.4511 0.0323 putative receptor; induced in stationary phase<br />
ymdA b1044 1.3743 0.0405 conserved hypothetical protein<br />
ymdE b1028 1.2439 0.0093 putative malonyl-CoA:Acyl carrier protein transacylase<br />
ymfM b1148 1.3536 0.0483 hypothetical protein<br />
ymgD b1171 0.4644 0.0021 hypothetical protein<br />
ynaE b1375 0.2665 0.0186 hypothetical protein<br />
ynaI b1330 0.5285 0.0218 YnaI<br />
ynbD b1411 1.4997 0.0330 putative enzyme<br />
yncC b1450 0.6862 0.0367 hypothetical protein<br />
yncD b1451 1.5934 0.0332 Probable TonB-dependent receptor<br />
yncI b1458 0.1672 0.0266 conserved protein<br />
yneI b1525 3.7228 0.0368 putative aldehyde dehydrogenase<br />
yneK b1527 0.9040 0.0285 conserved protein<br />
ynfN b1551 0.2403 0.0063 hypothetical protein<br />
yohJ b2141 2.9847 0.0353 putative transmembrane protein<br />
yohN b2107 0.6686 0.0265 hypothetical protein<br />
ypfJ b2475 1.4122 0.0100 conserved protein<br />
ypjF b2646 1.6516 0.0435 member <strong>of</strong> the YeeV, YkfI, YpjF family <strong>of</strong> toxin proteins<br />
yqaC b2657 0.3850 0.0243 putative enzyme<br />
yqeC b2876 0.6850 0.0170 conserved protein<br />
yqeI b2847 0.3106 0.0215 putative sensory transducer<br />
yqeJ b2848 0.3761 0.0379 conserved hypothetical protein<br />
yqgA b2966 2.0839 0.0160 putative transport protein<br />
yrhA b3443 0.7276 0.0181 conserved hypothetical protein<br />
ytfR b4485 1.7393 0.0402 subunit <strong>of</strong> YtfQ/YtfR/YtfS/YtfT/YjfF ABC transporter<br />
ytfT b4230 1.5937 0.0012 YtfQ/YtfR/YtfS/YtfT/YjfF ABC transporter<br />
zraS b4003 2.0921 0.0305 ZraS-Phis<br />
Supplementary table – V: CSH50 wild-type (A) Vs CSH50 hns-205::Tn10 (B), Transition to<br />
stationary phase<br />
<strong>gene</strong> blattner ratio p-value Function<br />
A/B<br />
aaeB b3240 1.1872 0.0075 hypothetical protein<br />
aceE b0114 0.2646 0.0654 subunit <strong>of</strong> E1p component <strong>of</strong> pyruvate dehydrogenase complex<br />
acrD b2470 1.3042 0.1589 AcrD<br />
ade b3665 0.0725 0.0088 cryptic adenine deaminase monomer<br />
adiA b4117 0.9019 0.0328 Adi<br />
agaB b3138 1.1736 0.1822 EIIAga<br />
alsC b4086 1.2567 0.1129 YjcV<br />
ansB b2957 0.2992 0.0537 asparaginase II<br />
arpB_2 b1721 0.9159 0.1043 hypothetical protein<br />
artJ b0860 0.8070 0.0643 arginine ABC transporter<br />
b1367 b1367 1.2731 0.0856 hypothetical protein<br />
b1567 b1567 0.7688 0.1747 hypothetical protein<br />
b3808 b3808 1.2642 0.0697 hypothetical protein<br />
b4103 b4103 1.4344 0.0818 putative C terminus <strong>of</strong> split phnE <strong>gene</strong> product<br />
betA b0311 1.2923 0.0890 choline dehydrogenase<br />
104