Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
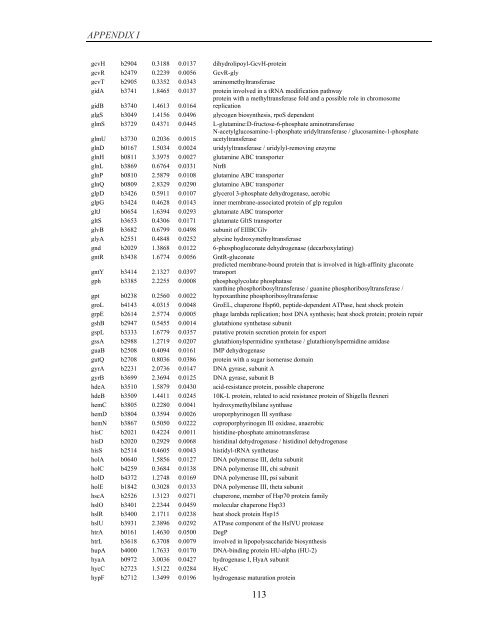
APPENDIX I<br />
gcvH b2904 0.3188 0.0137 dihydrolipoyl-GcvH-protein<br />
gcvR b2479 0.2239 0.0056 GcvR-gly<br />
gcvT b2905 0.3352 0.0343 aminomethyltransferase<br />
gidA b3741 1.8465 0.0137 protein involved in a tRNA modification pathway<br />
gidB b3740 1.4613 0.0164<br />
protein with a methyltransferase fold and a possible role in chromosome<br />
replication<br />
glgS b3049 1.4156 0.0496 glycogen biosynthesis, rpoS dependent<br />
glmS b3729 0.4371 0.0445 L-glutamine:D-fructose-6-phosphate aminotransferase<br />
glmU b3730 0.2036 0.0015<br />
N-acetylglucosamine-1-phosphate uridyltransferase / glucosamine-1-phosphate<br />
acetyltransferase<br />
glnD b0167 1.5034 0.0024 uridylyltransferase / uridylyl-removing enzyme<br />
glnH b0811 3.3975 0.0027 glutamine ABC transporter<br />
glnL b3869 0.6764 0.0331 NtrB<br />
glnP b0810 2.5879 0.0108 glutamine ABC transporter<br />
glnQ b0809 2.8329 0.0290 glutamine ABC transporter<br />
glpD b3426 0.5911 0.0107 glycerol 3-phosphate dehydrogenase, aerobic<br />
glpG b3424 0.4628 0.0143 inner membrane-associated protein <strong>of</strong> glp regulon<br />
gltJ b0654 1.6394 0.0293 glutamate ABC transporter<br />
gltS b3653 0.4306 0.0171 glutamate GltS transporter<br />
glvB b3682 0.6799 0.0498 subunit <strong>of</strong> EIIBCGlv<br />
glyA b2551 0.4848 0.0252 glycine hydroxymethyltransferase<br />
gnd b2029 1.3868 0.0122 6-phosphogluconate dehydrogenase (decarboxylating)<br />
gntR b3438 1.6774 0.0056 GntR-gluconate<br />
gntY b3414 2.1327 0.0397<br />
predicted membrane-bound protein that is involved in high-affinity gluconate<br />
transport<br />
gph b3385 2.2255 0.0008 phosphoglycolate phosphatase<br />
gpt b0238 0.2560 0.0022<br />
xanthine phosphoribosyltransferase / guanine phosphoribosyltransferase /<br />
hypoxanthine phosphoribosyltransferase<br />
groL b4143 4.0315 0.0048 GroEL, chaperone Hsp60, peptide-dependent ATPase, heat shock protein<br />
grpE b2614 2.5774 0.0005 phage lambda replication; host DNA synthesis; heat shock protein; protein repair<br />
gshB b2947 0.5455 0.0014 glutathione synthetase subunit<br />
gspL b3333 1.6779 0.0357 putative protein secretion protein for export<br />
gssA b2988 1.2719 0.0207 glutathionylspermidine synthetase / glutathionylspermidine amidase<br />
guaB b2508 0.4094 0.0161 IMP dehydrogenase<br />
gutQ b2708 0.8036 0.0386 protein with a sugar isomerase domain<br />
gyrA b2231 2.0736 0.0147 DNA gyrase, subunit A<br />
gyrB b3699 2.3694 0.0125 DNA gyrase, subunit B<br />
hdeA b3510 1.5879 0.0430 acid-resistance protein, possible chaperone<br />
hdeB b3509 1.4411 0.0245 10K-L protein, related to acid resistance protein <strong>of</strong> Shigella flexneri<br />
hemC b3805 0.2280 0.0041 hydroxymethylbilane synthase<br />
hemD b3804 0.3594 0.0026 uroporphyrinogen III synthase<br />
hemN b3867 0.5050 0.0222 coproporphyrinogen III oxidase, anaerobic<br />
hisC b2021 0.4224 0.0011 histidine-phosphate aminotransferase<br />
hisD b2020 0.2929 0.0068 histidinal dehydrogenase / histidinol dehydrogenase<br />
hisS b2514 0.4605 0.0043 histidyl-tRNA synthetase<br />
holA b0640 1.5856 0.0127 DNA polymerase III, delta subunit<br />
holC b4259 0.3684 0.0138 DNA polymerase III, chi subunit<br />
holD b4372 1.2748 0.0169 DNA polymerase III, psi subunit<br />
holE b1842 0.3028 0.0133 DNA polymerase III, theta subunit<br />
hscA b2526 1.3123 0.0271 chaperone, member <strong>of</strong> Hsp70 protein family<br />
hslO b3401 2.2344 0.0459 molecular chaperone Hsp33<br />
hslR b3400 2.1711 0.0238 heat shock protein Hsp15<br />
hslU b3931 2.3896 0.0292 ATPase component <strong>of</strong> the HslVU protease<br />
htrA b0161 1.4630 0.0500 DegP<br />
htrL b3618 6.3708 0.0079 involved in lipopolysaccharide biosynthesis<br />
hupA b4000 1.7633 0.0170 DNA-binding protein HU-alpha (HU-2)<br />
hyaA b0972 3.0036 0.0427 hydrogenase I, HyaA subunit<br />
hycC b2723 1.5122 0.0284 HycC<br />
hypF b2712 1.3499 0.0196 hydrogenase maturation protein<br />
113