Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
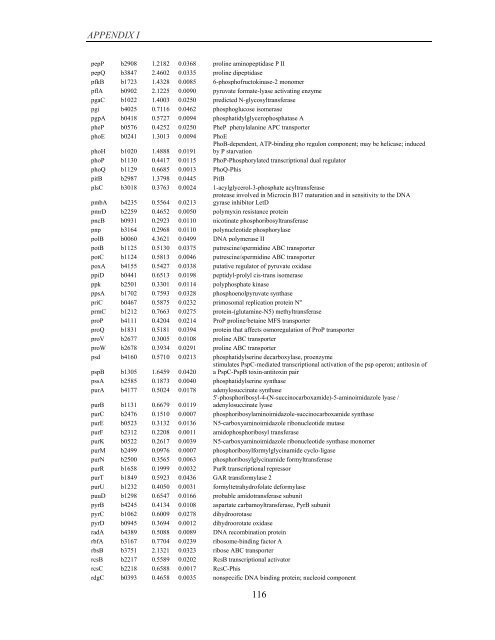
APPENDIX I<br />
pepP b2908 1.2182 0.0368 proline aminopeptidase P II<br />
pepQ b3847 2.4602 0.0335 proline dipeptidase<br />
pfkB b1723 1.4328 0.0085 6-phosph<strong>of</strong>ructokinase-2 monomer<br />
pflA b0902 2.1225 0.0090 pyruvate formate-lyase activating enzyme<br />
pgaC b1022 1.4003 0.0250 predicted N-glycosyltransferase<br />
pgi b4025 0.7116 0.0462 phosphoglucose isomerase<br />
pgpA b0418 0.5727 0.0094 phosphatidylglycerophosphatase A<br />
pheP b0576 0.4252 0.0250 PheP phenylalanine APC transporter<br />
phoE b0241 1.3013 0.0094 PhoE<br />
phoH b1020 1.4888 0.0191<br />
PhoB-dependent, ATP-binding pho regulon component; may be helicase; induced<br />
<strong>by</strong> P starvation<br />
phoP b1130 0.4417 0.0115 PhoP-Phosphorylated transcriptional dual regulator<br />
phoQ b1129 0.6685 0.0013 PhoQ-Phis<br />
pitB b2987 1.3798 0.0445 PitB<br />
plsC b3018 0.3763 0.0024 1-acylglycerol-3-phosphate acyltransferase<br />
pmbA b4235 0.5564 0.0213<br />
protease involved in Microcin B17 maturation and in sensitivity to the DNA<br />
gyrase inhibitor LetD<br />
pmrD b2259 0.4652 0.0050 polymyxin resistance protein<br />
pncB b0931 0.2923 0.0110 nicotinate phosphoribosyltransferase<br />
pnp b3164 0.2968 0.0110 polynucleotide phosphorylase<br />
polB b0060 4.3621 0.0499 DNA polymerase II<br />
potB b1125 0.5130 0.0375 putrescine/spermidine ABC transporter<br />
potC b1124 0.5813 0.0046 putrescine/spermidine ABC transporter<br />
poxA b4155 0.5427 0.0338 putative regulator <strong>of</strong> pyruvate oxidase<br />
ppiD b0441 0.6513 0.0198 peptidyl-prolyl cis-trans isomerase<br />
ppk b2501 0.3301 0.0114 polyphosphate kinase<br />
ppsA b1702 0.7593 0.0328 phosphoenolpyruvate synthase<br />
priC b0467 0.5875 0.0232 primosomal replication protein N''<br />
prmC b1212 0.7663 0.0275 protein-(glutamine-N5) methyltransferase<br />
proP b4111 0.4204 0.0214 ProP proline/betaine MFS transporter<br />
proQ b1831 0.5181 0.0394 protein that affects osmo<strong>regulation</strong> <strong>of</strong> ProP transporter<br />
proV b2677 0.3005 0.0108 proline ABC transporter<br />
proW b2678 0.3934 0.0291 proline ABC transporter<br />
psd b4160 0.5710 0.0213 phosphatidylserine decarboxylase, proenzyme<br />
pspB b1305 1.6459 0.0420<br />
stimulates PspC-mediated transcriptional activation <strong>of</strong> the psp operon; antitoxin <strong>of</strong><br />
a PspC-PspB toxin-antitoxin pair<br />
pssA b2585 0.1873 0.0040 phosphatidylserine synthase<br />
purA b4177 0.5024 0.0178 adenylosuccinate synthase<br />
purB b1131 0.6679 0.0119<br />
5'-phosphoribosyl-4-(N-succinocarboxamide)-5-aminoimidazole lyase /<br />
adenylosuccinate lyase<br />
purC b2476 0.1510 0.0007 phosphoribosylaminoimidazole-succinocarboxamide synthase<br />
purE b0523 0.3132 0.0136 N5-carboxyaminoimidazole ribonucleotide mutase<br />
purF b2312 0.2208 0.0011 amidophosphoribosyl transferase<br />
purK b0522 0.2617 0.0039 N5-carboxyaminoimidazole ribonucleotide synthase monomer<br />
purM b2499 0.0976 0.0007 phosphoribosylformylglycinamide cyclo-ligase<br />
purN b2500 0.3565 0.0063 phosphoribosylglycinamide formyltransferase<br />
purR b1658 0.1999 0.0032 PurR transcriptional repressor<br />
purT b1849 0.5923 0.0436 GAR transformylase 2<br />
purU b1232 0.4050 0.0031 formyltetrahydr<strong>of</strong>olate deformylase<br />
puuD b1298 0.6547 0.0166 probable amidotransferase subunit<br />
pyrB b4245 0.4134 0.0108 aspartate carbamoyltransferase, PyrB subunit<br />
pyrC b1062 0.6009 0.0278 dihydroorotase<br />
pyrD b0945 0.3694 0.0012 dihydroorotate oxidase<br />
radA b4389 0.5088 0.0089 DNA recombination protein<br />
rbfA b3167 0.7704 0.0239 ribosome-binding factor A<br />
rbsB b3751 2.1321 0.0323 ribose ABC transporter<br />
rcsB b2217 0.5589 0.0202 RcsB transcriptional activator<br />
rcsC b2218 0.6588 0.0017 RcsC-Phis<br />
rdgC b0393 0.4658 0.0035 nonspecific DNA binding protein; nucleoid component<br />
116