Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
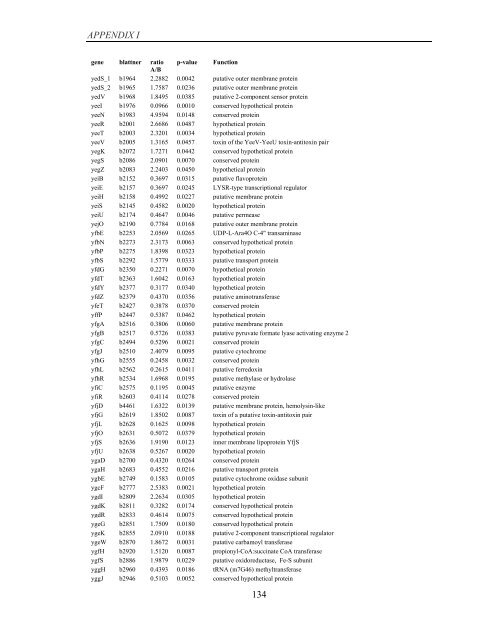
APPENDIX I<br />
<strong>gene</strong> blattner ratio p-value Function<br />
A/B<br />
yedS_1 b1964 2.2882 0.0042 putative outer membrane protein<br />
yedS_2 b1965 1.7587 0.0236 putative outer membrane protein<br />
yedV b1968 1.8495 0.0385 putative 2-component sensor protein<br />
yeeI b1976 0.0966 0.0010 conserved hypothetical protein<br />
yeeN b1983 4.9594 0.0148 conserved protein<br />
yeeR b2001 2.6686 0.0487 hypothetical protein<br />
yeeT b2003 2.3201 0.0034 hypothetical protein<br />
yeeV b2005 1.3165 0.0457 toxin <strong>of</strong> the YeeV-YeeU toxin-antitoxin pair<br />
yegK b2072 1.7271 0.0442 conserved hypothetical protein<br />
yegS b2086 2.0901 0.0070 conserved protein<br />
yegZ b2083 2.2403 0.0450 hypothetical protein<br />
yeiB b2152 0.3697 0.0315 putative flavoprotein<br />
yeiE b2157 0.3697 0.0245 LYSR-type transcriptional regulator<br />
yeiH b2158 0.4992 0.0227 putative membrane protein<br />
yeiS b2145 0.4582 0.0020 hypothetical protein<br />
yeiU b2174 0.4647 0.0046 putative permease<br />
yejO b2190 0.7784 0.0168 putative outer membrane protein<br />
yfbE b2253 2.0569 0.0265 UDP-L-Ara4O C-4" transaminase<br />
yfbN b2273 2.3173 0.0063 conserved hypothetical protein<br />
yfbP b2275 1.8398 0.0323 hypothetical protein<br />
yfbS b2292 1.5779 0.0333 putative transport protein<br />
yfdG b2350 0.2271 0.0070 hypothetical protein<br />
yfdT b2363 1.6042 0.0163 hypothetical protein<br />
yfdY b2377 0.3177 0.0340 hypothetical protein<br />
yfdZ b2379 0.4370 0.0356 putative aminotransferase<br />
yfeT b2427 0.3878 0.0370 conserved protein<br />
yffP b2447 0.5387 0.0462 hypothetical protein<br />
yfgA b2516 0.3806 0.0060 putative membrane protein<br />
yfgB b2517 0.5726 0.0383 putative pyruvate formate lyase activating enzyme 2<br />
yfgC b2494 0.5296 0.0021 conserved protein<br />
yfgJ b2510 2.4079 0.0095 putative cytochrome<br />
yfhG b2555 0.2458 0.0032 conserved protein<br />
yfhL b2562 0.2615 0.0411 putative ferredoxin<br />
yfhR b2534 1.6968 0.0195 putative methylase or hydrolase<br />
yfiC b2575 0.1195 0.0045 putative enzyme<br />
yfiR b2603 0.4114 0.0278 conserved protein<br />
yfjD b4461 1.6322 0.0139 putative membrane protein, hemolysin-like<br />
yfjG b2619 1.8502 0.0087 toxin <strong>of</strong> a putative toxin-antitoxin pair<br />
yfjL b2628 0.1625 0.0098 hypothetical protein<br />
yfjO b2631 0.5072 0.0379 hypothetical protein<br />
yfjS b2636 1.9190 0.0123 inner membrane lipoprotein YfjS<br />
yfjU b2638 0.5267 0.0020 hypothetical protein<br />
ygaD b2700 0.4320 0.0264 conserved protein<br />
ygaH b2683 0.4552 0.0216 putative transport protein<br />
ygbE b2749 0.1583 0.0105 putative cytochrome oxidase subunit<br />
ygcF b2777 2.5383 0.0021 hypothetical protein<br />
ygdI b2809 2.2634 0.0305 hypothetical protein<br />
ygdK b2811 0.3282 0.0174 conserved hypothetical protein<br />
ygdR b2833 0.4614 0.0075 conserved hypothetical protein<br />
ygeG b2851 1.7509 0.0180 conserved hypothetical protein<br />
ygeK b2855 2.0910 0.0188 putative 2-component transcriptional regulator<br />
ygeW b2870 1.8672 0.0031 putative carbamoyl transferase<br />
ygfH b2920 1.5120 0.0087 propionyl-CoA:succinate CoA transferase<br />
ygfS b2886 1.9879 0.0229 putative oxidoreductase, Fe-S subunit<br />
yggH b2960 0.4393 0.0186 tRNA (m7G46) methyltransferase<br />
yggJ b2946 0.5103 0.0052 conserved hypothetical protein<br />
134