Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
Coordinated regulation of gene expression by E ... - Jacobs University
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
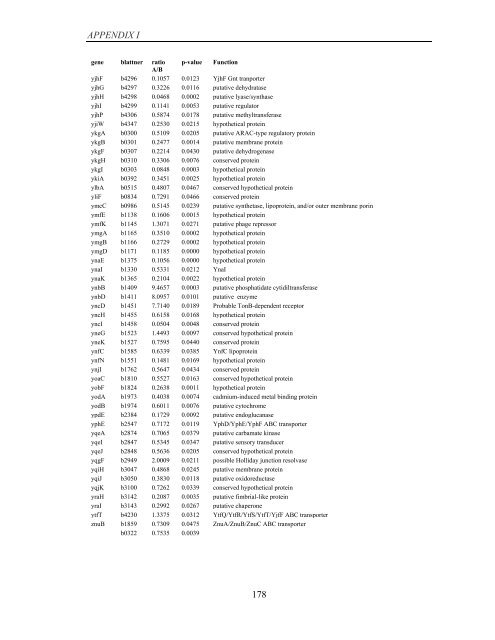
APPENDIX I<br />
<strong>gene</strong> blattner ratio p-value Function<br />
A/B<br />
yjhF b4296 0.1057 0.0123 YjhF Gnt tranporter<br />
yjhG b4297 0.3226 0.0116 putative dehydratase<br />
yjhH b4298 0.0468 0.0002 putative lyase/synthase<br />
yjhI b4299 0.1141 0.0053 putative regulator<br />
yjhP b4306 0.5874 0.0178 putative methyltransferase<br />
yjiW b4347 0.2530 0.0215 hypothetical protein<br />
ykgA b0300 0.5109 0.0205 putative ARAC-type regulatory protein<br />
ykgB b0301 0.2477 0.0014 putative membrane protein<br />
ykgF b0307 0.2214 0.0430 putative dehydrogenase<br />
ykgH b0310 0.3306 0.0076 conserved protein<br />
ykgI b0303 0.0848 0.0003 hypothetical protein<br />
ykiA b0392 0.3451 0.0025 hypothetical protein<br />
ylbA b0515 0.4807 0.0467 conserved hypothetical protein<br />
yliF b0834 0.7291 0.0466 conserved protein<br />
ymcC b0986 0.5145 0.0239 putative synthetase, lipoprotein, and/or outer membrane porin<br />
ymfE b1138 0.1606 0.0015 hypothetical protein<br />
ymfK b1145 1.3071 0.0271 putative phage repressor<br />
ymgA b1165 0.3510 0.0002 hypothetical protein<br />
ymgB b1166 0.2729 0.0002 hypothetical protein<br />
ymgD b1171 0.1185 0.0000 hypothetical protein<br />
ynaE b1375 0.1056 0.0000 hypothetical protein<br />
ynaI b1330 0.5331 0.0212 YnaI<br />
ynaK b1365 0.2104 0.0022 hypothetical protein<br />
ynbB b1409 9.4657 0.0003 putative phosphatidate cytidiltransferase<br />
ynbD b1411 8.0957 0.0101 putative enzyme<br />
yncD b1451 7.7140 0.0189 Probable TonB-dependent receptor<br />
yncH b1455 0.6158 0.0168 hypothetical protein<br />
yncI b1458 0.0504 0.0048 conserved protein<br />
yneG b1523 1.4493 0.0097 conserved hypothetical protein<br />
yneK b1527 0.7595 0.0440 conserved protein<br />
ynfC b1585 0.6339 0.0385 YnfC lipoprotein<br />
ynfN b1551 0.1481 0.0169 hypothetical protein<br />
ynjI b1762 0.5647 0.0434 conserved protein<br />
yoaC b1810 0.5527 0.0163 conserved hypothetical protein<br />
yobF b1824 0.2638 0.0011 hypothetical protein<br />
yodA b1973 0.4038 0.0074 cadmium-induced metal binding protein<br />
yodB b1974 0.6011 0.0076 putative cytochrome<br />
ypdE b2384 0.1729 0.0092 putative endoglucanase<br />
yphE b2547 0.7172 0.0119 YphD/YphE/YphF ABC transporter<br />
yqeA b2874 0.7065 0.0379 putative carbamate kinase<br />
yqeI b2847 0.5345 0.0347 putative sensory transducer<br />
yqeJ b2848 0.5636 0.0205 conserved hypothetical protein<br />
yqgF b2949 2.0009 0.0211 possible Holliday junction resolvase<br />
yqiH b3047 0.4868 0.0245 putative membrane protein<br />
yqiJ b3050 0.3830 0.0118 putative oxidoreductase<br />
yqjK b3100 0.7262 0.0339 conserved hypothetical protein<br />
yraH b3142 0.2087 0.0035 putative fimbrial-like protein<br />
yraI b3143 0.2992 0.0267 putative chaperone<br />
ytfT b4230 1.3375 0.0312 YtfQ/YtfR/YtfS/YtfT/YjfF ABC transporter<br />
znuB b1859 0.7309 0.0475 ZnuA/ZnuB/ZnuC ABC transporter<br />
b0322 0.7535 0.0039<br />
178