Zeitschrift für Rheumatologie – Supplement 1 - Deutsche ...
Zeitschrift für Rheumatologie – Supplement 1 - Deutsche ...
Zeitschrift für Rheumatologie – Supplement 1 - Deutsche ...
Erfolgreiche ePaper selbst erstellen
Machen Sie aus Ihren PDF Publikationen ein blätterbares Flipbook mit unserer einzigartigen Google optimierten e-Paper Software.
S18<br />
Abstracts<br />
FVFER1 8. Forum Experimentelle <strong>Rheumatologie</strong><br />
Teil 1: Genetics of Autoimmunity<br />
FVFER1-1<br />
COMP-assoziierte Chondrodysplasien sind keine Speicherkrankheit<br />
des endoplasmatischen Retikulums<br />
Dinser R. 1 , Weirich C. 1 , Schmitz A. 2 , M U. 1 , Paulsson M. 2 , Zaucke F. 2<br />
1 Klinik <strong>für</strong> Innere Medizin und <strong>Rheumatologie</strong>, Justus-Liebig Universität<br />
Gießen, Kerckhoff -Klinik, 2 Zentrum <strong>für</strong> Biochemie, Medizinische Fakultät,<br />
Universität zu Köln<br />
Einleitung: Chondrodysplasien mit Assoziation zu Mutationen des<br />
Cartilage Oligomeric Matrix Protein (COMP) sind durch disproportionierten<br />
Minderwuchs, schwere früh einsetzende Arthrose und Instabilität<br />
von Bändern und Sehnen charakterisiert. Bisher wurde die Pseudoachondroplasie<br />
(PSACH) als schwerste Form als Speicherkrankheit<br />
des endoplasmatischen Retikulums aufgefasst, da mutiertes COMP in<br />
lamellären Einschlüssen des endoplasmatischen Retikulums von Knorpelzellen<br />
aus Patientenbiopsien akkumuliert. Um die Pathophysiologie<br />
dieser Erkrankung zu untersuchen, haben wir ein Zellkulturmodell<br />
mittels adenoviralem Gentransfer entwickelt.<br />
Methodik: Wir verglichen drei PSACH-assoziierte COMP Mutationen,<br />
D475N, D469d, H587R durch Expression in primären Chondrozyten<br />
und Tenozyten mittels adenoviraler Transduktion. Für Langzeitanalysen<br />
wurden Chondrozyten in Alginat eingebettet.<br />
Ergebnisse: Mutation H587R wird vergleichbar zum wildtyp-Protein<br />
sezerniert, während die Mutationen D475N und D469d wie erwartet<br />
im endoplasmatischen Retikulum akkumulieren. Alle drei Mutationen<br />
verursachen schwere Fehlbildungen der in Kultur geformten Matrix<br />
sowie apoptotischen Zelltod unabhängig von ihrem Sekretionsverhalten<br />
und der untersuchten Zellart. Die Mutationen D475N und D469d<br />
induzieren eine „unfolded protein response“ im Gegensatz zu H587R.<br />
Schlussfolgerung: PSACH ist mehr als eine Speicherkrankheit des endoplasmatischen<br />
Retikulum, da selbst unverzögert sezernierte COMP-<br />
Mutationen eine Störung der Extrazellulärmatrix von Chondrozyten<br />
und Tenozyten verursachen sowie apoptotischen Zelltod induzieren.<br />
Vermutlich ist daher der Hauptfaktor der Pathogenese ein dominant negativer<br />
Eff ekt von mutiertem COMP auf die Kollagen-Fibrillogenese.<br />
FVFER1-2<br />
Collagen type II recognition by an arthritogenic murine autoantibody:<br />
no role of somatic mutation for high affi nity binding<br />
Sehnert B. 2 , Böiers U. 2 , Päßler S. 2 , Lanig H. 3 , Holmdahl R. 4 , Burkhardt1 1 Abteilung <strong>Rheumatologie</strong>, Johann Wolfgang Goethe-Universität Frankfurt<br />
am Main, 2 Medizinische Klinik III, Universität Erlangen<strong>–</strong>Nürnberg,<br />
3 4 Computer-Chemie-Zentrum, Universität Erlangen<strong>–</strong>Nürnberg, Lund<br />
University, Sweden<br />
Objectives: Th e aim of the present study was to characterize the interaction<br />
sites between the prototypic arthritogenic murine IgG mAB<br />
CIIC1that is highly somatically mutated and its epitope on type II collagen<br />
(CII, aa 359-369).<br />
Methods: Th e establishment of a dynamic simulation modelling of a<br />
CIIC1 single-chain fragment (scFV) in complex with the triple helical<br />
CIIC1 epitope permitted structural insights into immune complex formation.<br />
Th e computer-based data were experimentally tested by mutations<br />
of predicted critical residues into alanine in the C1scFvs and<br />
the respective CIIC1 epitope that were produced as recombinant constructs.<br />
Th e binding affi nities of the mutated scFVs were determined by<br />
ELISA and surface plasmon resonance measurements.<br />
Results: Th e mutation experiments confi rmed the predicted interaction<br />
sites of CII in the CDR2 heavy chain and CDR3 regions of both<br />
heavy and light chain. Surprinsingly also the model prediction, that the<br />
conversion of the C1scFv sequence into the respective germline does<br />
| <strong>Zeitschrift</strong> <strong>für</strong> <strong>Rheumatologie</strong> · <strong>Supplement</strong> 1 · 2006<br />
not aff ect CII binding affi nity (KD 3x 10-8) could be confi rmed experimentally<br />
by the mutagenesis of 13(!) positions.<br />
Conclusions: Our data indicate that potentially harmful cartilage specifi<br />
c humoral autoimmunity is germline encoded. Th e molecular modeling<br />
further demonstrate that the rigid collagen triple helix restricts<br />
the likelihood of molecular interactions with the corresponding CDR<br />
regions of the antibody considerably compared to globular antigens.<br />
Th ese sterical constraints might provide an explanation why somatic<br />
mutations have no obvious impact on CII recognition by the arthritogenic<br />
autoantibody. Moreover, the structural insights into CII-autoantibody<br />
interaction might be useful in future developments of collagenomimetic<br />
ligands for therapeutic and diagnosistic purposes.<br />
FVFER1-3<br />
A polymorphism in the 3´UTR of the TNFRII gene is associated with<br />
rheumatoid arthritis<br />
Schulz A., Pierer M., Kaltenhäuser S., Arnold S., Seidel W., Häntzschel H.,<br />
Wagner U.<br />
Department of Medicine IV, University of Leipzig<br />
Th e gene for tumor necrosis factor receptor II (TNFRII) is located on<br />
chromoson 1p36.2, a region, which has already been described to be<br />
associated with autoimmune diseases.<br />
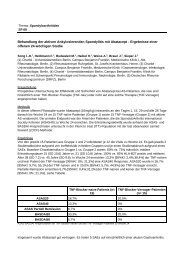
In the study presented here, a possible association of the following six<br />
TNFRII polymorphisms with rheumatoid arthritis (RA) was analyzed:<br />
two polymorphisms in exon 6 (gb:M32315: 676 T/G, 783 G/A), which<br />
result in amino acid changes (Met196Arg and Glu232Lys); one polymorphism<br />
in exon 2 (257 A/G); and three polymorphism in exon 10<br />
(1663 G/A, 1668 T/G, 1690 T/C), which are located in the 3´UTR.<br />
To identify these polymorphisms, polymerase chain reaction based<br />
assays with sequence specifi c primers were used. Genomic DNA of<br />
358 patients with RA according to the 1987 ACR criteria and 351 healthy<br />
controls was genotyped.<br />
Th e 1668 G allele in the 3´UTR (exon 10) was found more frequently in<br />
RA patients than in healthy controls (5.45% vs. 2.71%, odds ratio= 2.07,<br />
p=0.022).<br />
For the 676 T/G polymorphism in exon 6, only a non-signifi cant trend<br />
towards a higher frequency in RA patients was found. All other examined<br />
polymorphisms were not associated with RA.<br />
Th e results suggest that a polymorphism in the 3´UTR of the TNFRII<br />
gene is associated with RA.<br />
FVFER1-4<br />
Analysis of mutations/SNPs in jun and fos proto-oncogene promoters<br />
in the rheumatoid arthritis synovial membrane<br />
Huber R. 1 , Lanick B. 1 , Ukena B. 1 , Dunger S. 2 , Pohlers D. 1 , Kinne RW. 1<br />
1 Experimental Rheumatology Unit, Department of Orthopedics, Friedrich-<br />
Schiller-University Jena, Waldkrankenhaus, 2 Franz-Volhard Clinic, Department<br />
of Cardiology, HELIOS Klinikum-Berlin, Charité Campus Berlin-Buch<br />
Objective: To identify relevant mutations and single nucleotide polymorphisms<br />
(SNPs) in the promoters of jun and fos proto-oncogenes<br />
in the synovial membrane (SM) of rheumatoid arthritis (RA), osteoarthritis<br />
(OA), and normal control (NC) samples.<br />
Methods: Mutations/SNPs in the respective jun/fos promoters were<br />
identifi ed in DNA samples isolated from RA (n=10), OA (n=10), and<br />
NC (n=5) SM tissue and blood by Non-Isotopic RNase Cleavage Assay.<br />
Wildtype (wt) and mutated promoter fragments were cloned into<br />
the reporter gene expression vector pUBT-luc (expressing luciferase).<br />
Reporter gene expression of the respective promoter fragments was<br />
measured in NIH 3T3 cells one and two days aft er transfection (both<br />
constitutively and following stimulation with PMA) using a dual luciferase<br />
assay system.