ABI Prism® 7900HT Sequence Detection System ... - OpenWetWare
ABI Prism® 7900HT Sequence Detection System ... - OpenWetWare
ABI Prism® 7900HT Sequence Detection System ... - OpenWetWare
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
Preparing and<br />
Running an<br />
RNase P Plate<br />
Verifying<br />
Instrument<br />
Performance<br />
7-26 <strong>System</strong> Maintenance<br />
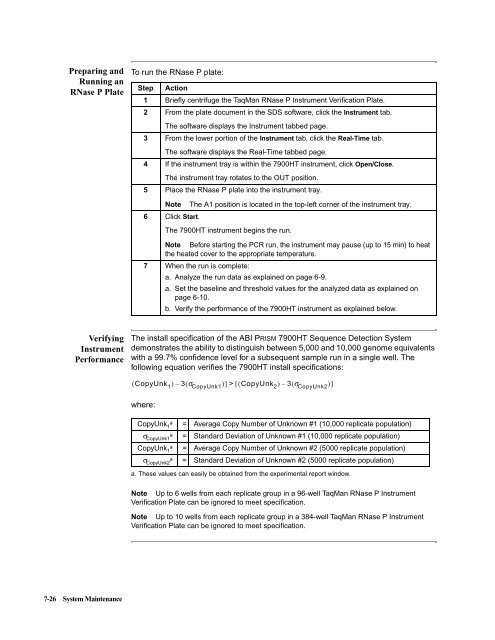
To run the RNase P plate:<br />
Step Action<br />
1 Briefly centrifuge the TaqMan RNase P Instrument Verification Plate.<br />
2 From the plate document in the SDS software, click the Instrument tab.<br />
The software displays the Instrument tabbed page.<br />
3 From the lower portion of the Instrument tab, click the Real-Time tab.<br />
The software displays the Real-Time tabbed page.<br />
4 If the instrument tray is within the <strong>7900HT</strong> instrument, click Open/Close.<br />
The instrument tray rotates to the OUT position.<br />
5 Place the RNase P plate into the instrument tray.<br />
Note The A1 position is located in the top-left corner of the instrument tray.<br />
6 Click Start.<br />
The install specification of the <strong>ABI</strong> PRISM <strong>7900HT</strong> <strong>Sequence</strong> <strong>Detection</strong> <strong>System</strong><br />
demonstrates the ability to distinguish between 5,000 and 10,000 genome equivalents<br />
with a 99.7% confidence level for a subsequent sample run in a single well. The<br />
following equation verifies the <strong>7900HT</strong> install specifications:<br />
where:<br />
The <strong>7900HT</strong> instrument begins the run.<br />
Note Before starting the PCR run, the instrument may pause (up to 15 min) to heat<br />
the heated cover to the appropriate temperature.<br />
7 When the run is complete:<br />
a. Analyze the run data as explained on page 6-9.<br />
a. Set the baseline and threshold values for the analyzed data as explained on<br />
page 6-10.<br />
b. Verify the performance of the <strong>7900HT</strong> instrument as explained below.<br />
[ ( CopyUnk1) – 3( σCopyUnk1) ] ><br />
CopyUnk2 [ ( ) – 3( σCopyUnk2) ]<br />
CopyUnk a<br />
1 = Average Copy Number of Unknown #1 (10,000 replicate population)<br />
σ a<br />
CopyUnk1 = Standard Deviation of Unknown #1 (10,000 replicate population)<br />
CopyUnk a<br />
1 = Average Copy Number of Unknown #2 (5000 replicate population)<br />
σ a<br />
CopyUnk2 = Standard Deviation of Unknown #2 (5000 replicate population)<br />
a. These values can easily be obtained from the experimental report window.<br />
Note Up to 6 wells from each replicate group in a 96-well TaqMan RNase P Instrument<br />
Verification Plate can be ignored to meet specification.<br />
Note Up to 10 wells from each replicate group in a 384-well TaqMan RNase P Instrument<br />
Verification Plate can be ignored to meet specification.